- Services
- FAQ
- Case Study
- Demo
- Related Services
- Support Documents
- Inquiry
Dynamic Light Scattering (DLS) Services for Biopharmaceutical Development
At Creative Proteomics, our Dynamic Light Scattering (DLS) services are strategically designed to support particle size determination, protein aggregation analysis, and antibody drug development workflows across early discovery, pre-formulation, and CMC development stages. With rich experience in biophysical characterization, we deliver high-sensitivity DLS measurements tailored to the real-world challenges faced by biomedical researchers, pharmaceutical R&D teams, and CRO collaborators.
DLS plays a central role in evaluating hydrodynamic size distributions, colloidal stability, and aggregation behavior of therapeutic proteins, monoclonal antibodies, viral vectors, and nanomedicines. Our service platform integrates DLS with static light scattering (SLS), zeta potential analysis, and orthogonal chromatographic methods to deliver robust, decision-enabling data outputs.
For clients requiring comprehensive characterization, DLS integrates seamlessly with our:
- Protein Aggregation Analysis Services
- Antibody Drug Characterization Services
- Biopharmaceutical Stability Testing Services
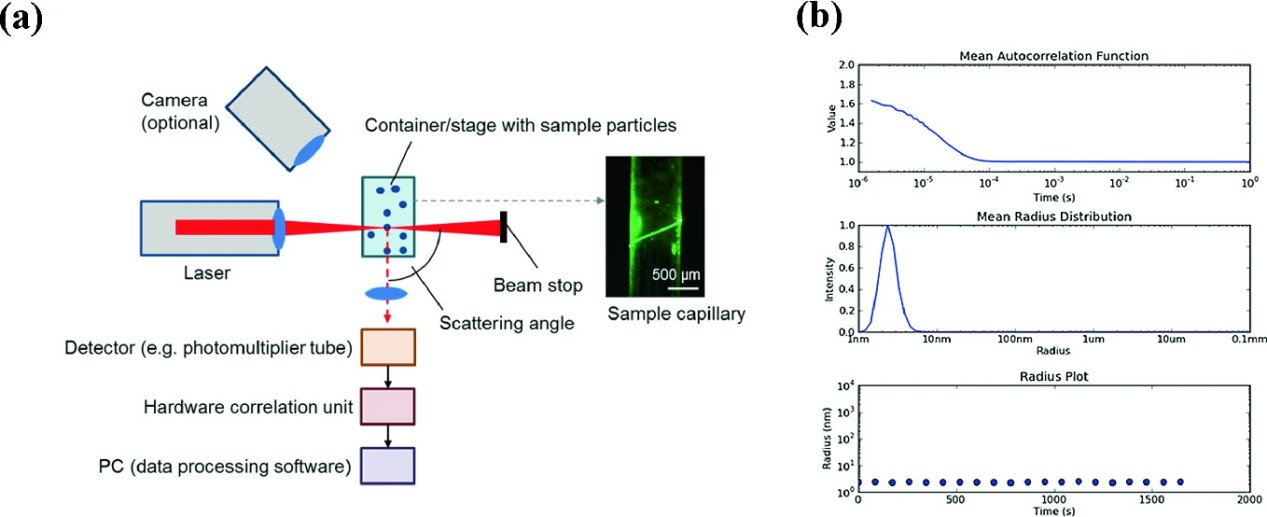 Figure 1. Scheme of a light scattering setup (Falke S, et al., 2019).
Figure 1. Scheme of a light scattering setup (Falke S, et al., 2019).Advantages of Our DLS Services
- High sensitivity to early aggregation events
- Broad particle size measurement range
- Minimal sample preparation
- Low sample consumption
- High-throughput screening capability
- Temperature-controlled stability profiling
Our DLS services are fully customizable and seamlessly integrated into broader proteomics and stability testing workflows.
Protein Aggregation Analysis in Antibody Drug Development
Protein aggregation remains one of the most significant risks in antibody drug development. Even low levels of soluble aggregates can compromise stability, reduce efficacy, and complicate manufacturing.
Our DLS-based protein aggregation analysis enables:
- Early detection of oligomers and pre-aggregates
- Quantification of aggregate size distribution
- Comparative stability profiling under stress conditions
- High-throughput formulation screening
For advanced aggregate profiling, we integrate DLS with complementary techniques, such as SEC-MALS, to obtain absolute molecular weight confirmation and improve resolution in heterogeneous systems.
Advanced Particle Size Determination for Biologics
Our DLS services support size characterization across:
- Monoclonal antibodies and antibody fragments
- Fusion proteins and bispecific constructs
- Viral vectors (AAV, lentivirus)
- Lipid nanoparticles (LNPs) and liposomes
- Protein-based nanocapsules
- Polymer conjugates and macromolecular complexes
Protein-Protein Interaction Studies and Colloidal Stability Assessment
Beyond size measurement, DLS provides quantitative insight into intermolecular interactions that drive aggregation behavior.
We determine:
- Diffusion interaction parameter (kD)
- Second virial coefficient (B22)
- Theoretical diffusion constant at zero concentration (D0)
Thermal Stability and Stress-Induced Aggregation Profiling
Temperature-induced stress studies are indispensable for evaluating antibody stability and predicting long-term behavior.
Our DLS platform supports:
- Temperature ramp experiments (4°C–85°C)
- Determinant of aggregation onset temperature
- Monitoring of Tsize and Tscattering
- Thermal denaturation curve analysis
Integrated DLS Platforms
Our instrumentation supports both cuvette-based precision analysis and high-throughput plate formats (96-1536 wells), enabling:
- Design-of-Experiment (DoE) formulation screening
- Parallel condition comparison
- Minimal sample volume (as low as 2-4 µL)
Characterization of Nanomedicines and Gene Therapy Vectors
In addition to protein therapeutics, DLS is widely applied to advanced delivery systems, including:
- Lipid nanoparticles (LNPs)
- Liposomes
- Virus-like particles
- Adeno-associated virus (AAV) preparations
- Polymers-based nanocarriers
DLS provides rapid evaluation of:
- Particle size distribution
- Aggregate content
- Particle concentration (when combined with static light scattering)
- Thermal stability
Creative Proteomics' DLS Service Workflow
- Sample Preparation: Controlled handling to minimize dust/particulate contamination; optimize concentration, buffer, and viscosity.
- Experimental Design: Tailored measurement conditions; high-throughput plate-based screening for multiple formulations.
- DLS Measurement: Laser-based detection of Brownian motion; autocorrelation analysis to calculate hydrodynamic radius.
- Data Analysis: Z-average particle size, polydispersity index, intensity- and volume-weighted distributions; detect aggregates and oligomers.
- Stability & Interaction Assessment: Evaluate thermal stability, aggregation onset, and protein–protein interactions; derive diffusion and interaction parameters.
Sample Requirements
| Sample Type | Proteins, peptides, antibodies, oligonucleotides, nanoparticles, liposomes, lipid nanoparticles, virus-like particles |
| Concentration Range | 0.1 mg/mL – 100 mg/mL (protein samples) 10^9 – 10^12 particles/mL (nanoparticles/viral vectors) |
| Volume Required | 2–10 µL for plate-based assays 20–100 µL for cuvette-based measurements |
| Buffer Conditions | Typical formulation buffers are acceptable; avoid high particulate contamination |
| Temperature Requirements | 4°C – 85°C |
Why Choose Creative Proteomics for DLS Services?
- Customized method development
- Integrated stability and aggregation workflows
- Expert interpretation beyond raw size data
- Rapid turnaround times
- Comprehensive reporting tailored to regulatory and development needs
FAQ
Q1: How does Dynamic Light Scattering (DLS) measure particle size?
A1: DLS measures the size of particles in a solution by tracking their natural motion, called Brownian motion. A laser shines through the sample, and the scattered light fluctuates as particles move. These fluctuations are analyzed to calculate the particle's diffusion rate, which is then converted into a size estimate called the hydrodynamic radius. This method works for proteins, nanoparticles, and other biomolecules without requiring separation.
Q2: How does DLS detect protein–protein interactions?
A2: DLS can measure whether proteins attract or repel each other in solution. By analyzing how diffusion changes with concentration, we can quantify interaction tendencies. Proteins that stick together form larger complexes, which increase the measured particle size. This information helps optimize high-concentration formulations, such as antibody therapies.
Q3: What is the difference between DLS and other particle sizing methods?
A3:
| Method | Strength | Limitation |
|---|---|---|
| DLS | Rapid, solution-based, minimal sample prep | Sensitive to large aggregates |
| Size Exclusion Chromatography | Separates species | Requires longer analysis time |
| Electron Microscopy | Direct visualization | Complex preparation |
| Nanoparticle Tracking Analysis | Individual particle tracking | Requires optimal concentration range |
Q4: How does DLS complement other analytical methods?
A4: DLS is fast, non-destructive, and requires minimal sample preparation, making it ideal for early screening. For detailed structural insights or highly complex samples, techniques like size-exclusion chromatography, electron microscopy, or multi-angle light scattering can be used alongside DLS. Together, these methods provide a comprehensive view of particle size, aggregation, and stability.
Case Study
Case: Aggregation Kinetics for IgG1-Based Monoclonal Antibody Therapeutics
Authors: Singla A, et al.
Journal: The AAPS journal
Year: 2016
DOI: 10.1208/s12248-016-9887-0
Background
Monoclonal antibodies (mAbs) are widely used as therapeutic agents for diseases such as cancer, autoimmune disorders, and infectious diseases. However, protein aggregation is a major challenge in antibody drug development because it can lead to loss of biological activity, reduced product quality, and increased immunogenicity. Aggregation may occur during production, purification, formulation, or storage. Understanding the mechanisms and kinetics of aggregation under different environmental conditions is essential for improving antibody stability and manufacturing processes.
Methods
- Size Exclusion Chromatography (SEC): to measure monomer loss and quantify aggregate formation.
- Dynamic Light Scattering (DLS): to determine the hydrodynamic size of antibody aggregates and identify oligomer species such as monomers, dimers, trimers, tetramers, and pentamers.
- Circular Dichroism (CD) spectroscopy: to analyze structural changes in the antibody's secondary structure.
Results
The study found that environmental conditions strongly influence antibody aggregation. DLS analysis confirmed the formation of multiple oligomeric species, with dimers acting as the critical nucleus for aggregation.
Key findings include:
- pH: Low pH (pH 3.0) significantly increased aggregation, while higher pH values maintained stability.
- Temperature: Aggregation increased with temperature, with the highest aggregation observed at 30°C.
- Salt concentration: Increasing salt levels accelerated aggregation by reducing electrostatic repulsion between protein molecules.
- Buffer type: Citrate buffer promoted aggregation more than acetate or glycine buffers.
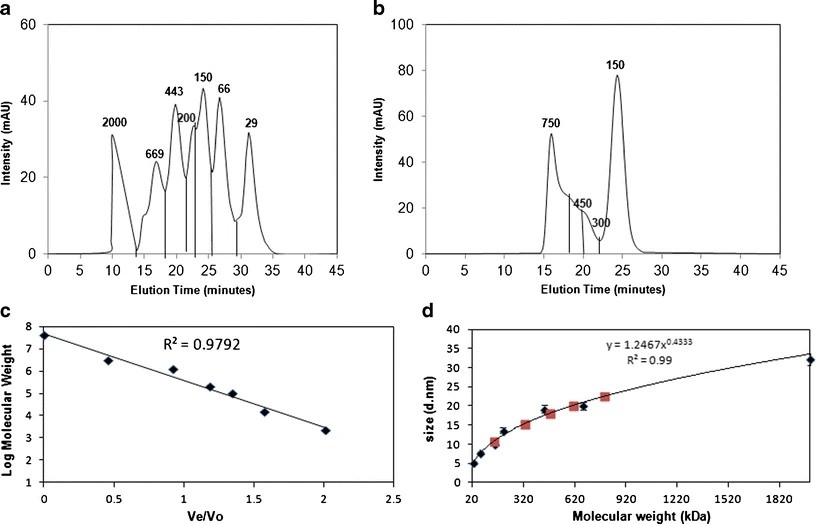 Figure 2. SEC chromatograms and DLS data for oligomer distribution analysis.
Figure 2. SEC chromatograms and DLS data for oligomer distribution analysis.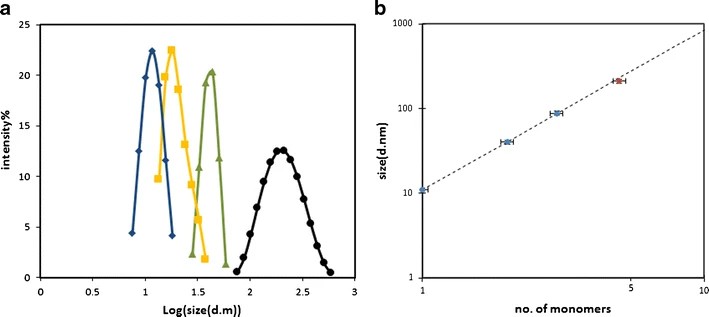 Figure 3. DLS data of various aggregate species separated by SEC.
Figure 3. DLS data of various aggregate species separated by SEC.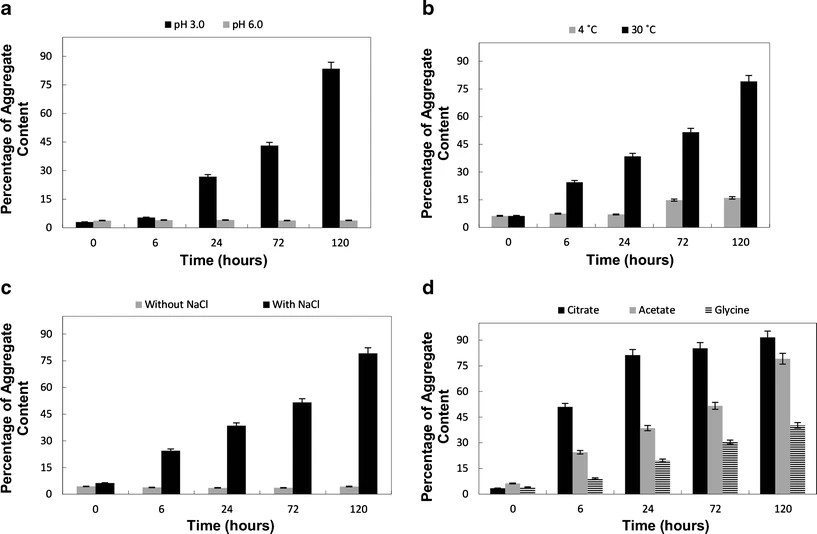 Figure 4. Aggregation of a monoclonal antibody therapeutic.
Figure 4. Aggregation of a monoclonal antibody therapeutic.Conclusion
The study concludes that aggregation of IgG1 monoclonal antibodies is strongly influenced by environmental conditions, particularly pH, temperature, salt concentration, and buffer composition. The results indicate that aggregation follows a nucleation-driven mechanism, with dimers serving as the initial nuclei for aggregate growth.
Demo
Demo: The characterization of exosomes from fibrosarcoma cell and the useful usage of Dynamic Light Scattering (DLS) for their evaluation
This study characterizes fibrosarcoma-derived exosomes and demonstrates that dynamic light scattering (DLS) enables rapid measurement of their size and homogeneity.
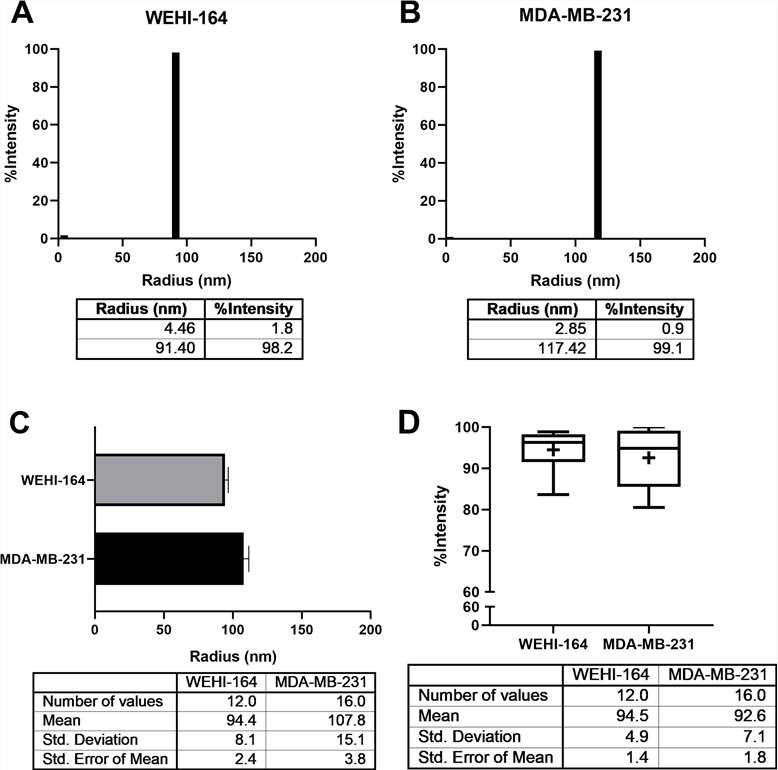 Figure 5. The average radius of exosomes derived from the WEHI-164 cell line was identified by DLS.
Figure 5. The average radius of exosomes derived from the WEHI-164 cell line was identified by DLS.References
- Singla A, et al. Aggregation kinetics for IgG1-based monoclonal antibody therapeutics. The AAPS journal, 2016, 18(3): 689-702. DOI: 10.1208/s12248-016-9887-0
- Falke S, Betzel C. Dynamic light scattering (DLS) principles, perspectives, applications to biological samples. Radiation in bioanalysis: Spectroscopic techniques and theoretical methods, 2019: 173-193. DOI: 10.1007/978-3-030-28247-9_6
- Jia Z, et al. Dynamic light scattering: a powerful tool for in situ nanoparticle sizing. Colloids and Interfaces, 2023, 7(1): 15. DOI: 10.3390/colloids7010015
- Lyu T S, et al. The characterization of exosomes from fibrosarcoma cell and the useful usage of Dynamic Light Scattering (DLS) for their evaluation. PLoS One, 2021, 16(1): e0231994. DOI: 10.1371/journal.pone.0231994
Related Services
Support Documents
KNOWLEDGE CENTER