The term "extinction coefficient" refers to the degree of light absorption by a measured solution. When the solution concentration is high, resulting in a darker color after chromogenic development, there is a pronounced absorption of light, leading to a decrease in light transmittance. Conversely, at lower concentrations with a lighter color, the absorption of light is diminished, resulting in higher light transmittance. For a given solution, it exhibits distinct absorption peaks for light of different wavelengths.
To enhance sensitivity, it is customary to select the complementary color of light as the preferred wavelength. For instance, blue and yellow are complementary colors, with 595nm wavelength falling within this range, yielding the maximum absorption value and thereby enhancing sensitivity. In contrast, 465nm corresponds to cyan light, and as blue solutions exhibit lower absorption at this wavelength, the sensitivity is relatively diminished.
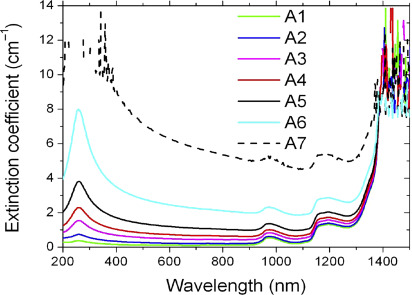 Spectral extinction coefficient (Preparation, Characterization, Properties and Application of Nanofluid, 2019)
Spectral extinction coefficient (Preparation, Characterization, Properties and Application of Nanofluid, 2019)Select Service
In numerous applications involving peptides or proteins, the identification of protein-containing fractions or the estimation of the concentration of purified samples is of paramount importance. Amino acids harboring aromatic side chains, namely tyrosine, tryptophan, and phenylalanine, exhibit strong ultraviolet (UV) light absorption. Consequently, the absorption of ultraviolet light by proteins and peptides is directly proportional to the content of their aromatic amino acids and the total concentration. Once the specific absorption coefficient for a given protein, determined by its fixed amino acid composition, is established, the protein concentration in a solution can be calculated from its absorbance.
For the majority of proteins, ultraviolet (UV) light absorption allows detection at concentrations as low as 100 µg/mL. However, in the case of complex protein solutions, such as cell lysates, estimating protein concentration through UV absorption is not precise due to the unclear composition of proteins with different absorption coefficients. Additionally, proteins are not the sole molecules capable of UV absorption; complex solutions often contain compounds like nucleic acids that can interfere with the determination of protein concentration using this method. Nevertheless, for commonly used protein aqueous solutions in research laboratory settings, interference from other compounds can be minimized by measuring absorbance at 280 nm.
Only tryptophan (Trp, W) and tyrosine (Tyr, Y), along with a lesser extent of cysteine (Cys, C), significantly contribute to the absorbance of peptides or proteins at 280 nm. Phenylalanine (Phe, F) exhibits absorption primarily at lower wavelengths (240-265 nm).
Absorbance and Extinction Coefficient
The ratio of the transmitted radiant power (P) through a sample to the radiant power incident upon the sample (P0) is termed transmittance (T):
T = P0/P
Consequently, absorbance (A) is defined as the logarithm (base 10) of the reciprocal of transmittance:
A −logT = logT1
In a spectrophotometer, monochromatic parallel light enters the sample perpendicularly, forming a straight line with the sample's plane. Under these conditions, the transmittance and absorbance of the sample depend on the molar concentration (c), path length (cm), and molar absorptivity (ε) of the dissolved substance at a specified wavelength (λ) [1].
Aλ=ε⋅ c⋅ L
Beer's law asserts that for a specific substance dissolved in a particular solvent, the molar absorptivity measured at a specific wavelength is constant (absorbance is directly proportional to concentration) [2]. Due to this, molar absorptivity is termed molar absorption coefficient or molar extinction coefficient. As transmittance and absorbance are dimensionless, the unit of molar absorptivity must cancel out with the units of concentration and path length measurements. Therefore, the unit of molar absorptivity is M−1cm−1. Standard laboratory spectrophotometers are designed for 1 cm width sample cuvettes; hence, the path length is often assumed to be 1 cm in most calculations.
Aλ= εcL = εc when L = 1cm
The molar absorptivity of peptides or proteins is related to their amino acid composition, specifically tryptophan (W), tyrosine (Y), and cysteine (C). At 280 nm, this value is approximated as the weighted sum of the molar absorptivities of these three amino acids, as expressed by the equation [3,4]:
ε=(nW×5500) + (nY×1490) + (nC×125)
Where n is the quantity of each residue, and the numerical values represent the molar absorptivities of the amino acids at 280 nm.
Determining Protein Concentration Based on Absorbance
In elucidating the concentration expression of Beer's Law, a comprehensive understanding emerges, providing insight into the requisite data for determining the concentration of peptide or protein solutions:
C = A/εL (or C = A/ ε when L = 1cm).
By dividing the measured absorbance of the peptide or protein solution by the calculated or known molar extinction coefficient, the molar concentration of the peptide or protein solution can be derived. To ensure precision in the calculation, the amino acid composition of the peptide or protein must be known, enabling the application of the aforementioned formula to compute the molar extinction coefficient.
For complex molecules such as peptides or proteins, a universal molar absorptivity value does not exist. Even minor variations in buffer type, ion strength, and pH can exert a subtle influence on absorbance values. In reality, most protein formulations, even with identical purity, exhibit differences in conformation and modification degrees, such as oxidation, all of which can impact absorbance. Therefore, the optimal molar absorptivity value is determined empirically by dissolving a known concentration of the research protein solution in the same buffer as the sample.
Furthermore, numerous absorptivity values (i.e., molar extinction coefficients) for proteins have been compiled from the literature. These values provide sufficient accuracy for the majority of routine laboratory applications requiring the assessment of protein concentration. Most data report protein molar absorptivity measured at or near the 280 nm wavelength in phosphate or other physiological buffers.
Application of Molar Absorptivity to 1% Solution Absorbance:
In computations, the utilization of molar absorptivity allows for the derivation of concentration expression in molar units:
A/ εmolar= Molar Concentration.
However, many sources, including the references mentioned earlier, do not provide molar absorptivity. Instead, they offer absorbance (A280nm) values measured in a 1 cm cuvette for a 1% (=1 g/100 mL) solution. These values are understood as a percentage molar absorptivity (εpercent), with units of (g/100 mL)-1cm-1 rather than 1M-1cm-1. Therefore, when these values are applied as absorptivity in a general formula, the concentration unit c should be in solution percentage (i.e., 1%=1 g/100 mL=10 mg/mL).
A/εpercent =Concentration Percentage
If reporting concentration in units of mg/mL is desired, an adjustment factor of 10 (i.e., conversion from 10 mg/mL to 1 mg/mL concentration units) must be applied when using these solution percentage absorptivity values.
(A/εpercent)×10=Concentration in mg/mL
The relationship between molar absorptivity (εpercent) and percentage absorptivity (εpercent) is as follows:
εpercent×10=εpercent×(Protein Molecular Weight)
Some data also provide absorbance values for 0.1% (=mg/mL) protein solutions, as this measurement unit is more convenient and common in protein work than percentage solutions. The variation in reporting highlights the importance of careful scrutiny of these values to ensure the understanding and accurate application of measurement units.
Example A: Proteins and Protein Mixtures with Unknown Extinction Coefficients
In cases where extinction coefficient information is lacking, a preliminary estimation of the protein concentration in protein or protein mixtures solutions can be made by assuming a value of 10 for εpercent. The extinction coefficients (εpercent) for most proteins typically fall within the range of 4.0 to 24.0 [5]. Therefore, even though specific proteins may have varying εpercent values, the average for a mixture of proteins could be approximated to be around 10.
Example B: Immunoglobulins
The protein extinction coefficient (ε) for most mammalian antibodies, known as immunoglobulins, typically falls within the range of 12 to 15. Therefore, for a typical antibody solution, let's assume
A1%280nm =14 or A1mg/ml280nm =14.
For a typical IgG with a molecular weight (MW) of 150,000, this value corresponds to a molar extinction coefficient (ε) of 210,000 M-1cm-1.
References
- Lange's Handbook of Chemistry, 14th Edition, Dean, J.A., Ed. (1992). McGraw-Hill, Inc., New York.
- Handbook of Chemistry and Physics, 56th Edition, Weast, R.C., Ed. (1975). CRC Press, Cleveland.
- Gill, S.C. and von Hippel, P.H. (1989). Calculation of protein extinction coefficients from amino acid sequence data. Anal. Biochem. 182:319-26.
- Pace, C.N., et al. (1995). How to measure and predict the molar absorption coefficient of a protein. Protein Sci. 4:2411-23.
- Practical Handbook of Biochemistry and Molecular Biology, Fasman, D.G., Ed. (1992). CRC Press, Boston.