Venom Genomics Analysis

Creative Proteomics has been providing researchers with accurate and affordable bioinformatics data analysis services for many years. For venom research, we provide customers with genomic biological information data analysis for various research purposes, such as de novo sequencing data analysis, genome resequencing data analysis, and exome sequencing data analysis, etc. The venom genome data analysis content we provide is applicable to various venomous species, such as snakes, scorpions, frogs, venomous lizards, fire-bellied toads, snails, spiders, bees, ants, cone snails, jellyfish, etc. More importantly, More importantly, with our years of project experience, we can analyze high-throughput sequencing data generated by different platforms, such as Illumina (short reading) and PacBio (long reading) platforms.
Introduction
Genome analysis is a technology to obtain the base sequence of target DNA and RNA fragments. With the development of next-generation sequencing technology and third-generation sequencing technology, massive amounts of genomic data has been generated. Information mining of target DNA and RNA fragment sequences obtained by bioinformatics methods is the basis for further molecular biology research. For venom genome analysis, Creative Proteomics provides the following analysis services:
For some toxic species without a reference genome, we provide de novo sequencing bioinformatics data analysis, which can assemble a certain toxic species without any reference sequence information, and use bioinformatics analysis methods to splice and assemble to obtain the species’ genomic sequence diagram.
Venom Genomic Data Analysis Pipeline
The venom genomics data analysis service provided by Creative Proteomics, from raw data processing and selection of software parameters, and then to the generation of result reports, has undergone scientific and meticulous design and strict quality control at every step to ensure high-quality research results. The general pipeline for venom de novo sequencing data analysis (an example of venom genomics data analysis) is outlined below.
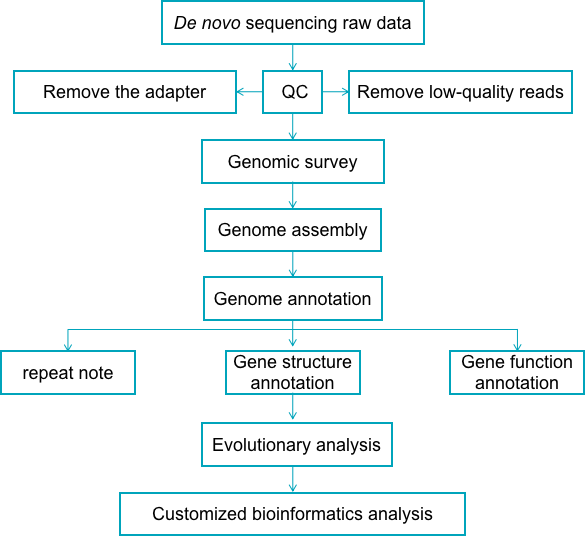 Fig 1. Venom de novo sequencing data analysis pipeline - Creative Proteomics
Fig 1. Venom de novo sequencing data analysis pipeline - Creative ProteomicsApplication of Venom Genomics Analysis
- For poisonous species without a reference genome, through the analysis of de novo sequencing data, the draft genome is drawn, and the reference sequence of the species is obtained, which provides a data basis for subsequent research on the species.
- Detection of various mutations: It can detect various mutation types such as SNP, InDel, SV and CNV, and can be used to explain the diversity of venom through gene or sequence polymorphism.
- Mining the variation information of venom-related functional genes, further studying the characteristics of poisonous species, population evolution issues, and locating target trait gene loci.
- Detect high-frequency, low-frequency, and even rare point mutations and structural variation information related to venom at the genome level.
- Build a poisonous species database.
Creative Proteomics, as a rare professional venomomics research service provider in the world, relies on its years of bioinformatics data analysis experience to provide customers with one-stop venom genomics data analysis services. From sample preparation, to high-throughput sequencing, to personalized bioinformatics services, we provide researchers with one-stop venom genomics research solutions, aiming to help researcher to analyze the venom synthesis mechanism, venom evolution, genetic variation, regulation and other issues from the genome level. If you have any questions about our venom genomics data analysis services, please feel free to contact our technical support, the technical support will reply to you as soon as possible. Look forward to working with you!
Reference
For research use only. Not intended for any clinical use.





 Fig 1. Venom de novo sequencing data analysis pipeline - Creative Proteomics
Fig 1. Venom de novo sequencing data analysis pipeline - Creative Proteomics