What Is Targeted Proteomics and Why Make the Switch?
While discovery proteomics (like DDA or DIA) is designed to cast a wide net and identify thousands of proteins, it often lacks the statistical power and absolute quantitative rigor needed for translational validation. Targeted proteomics flips the paradigm. Instead of scanning everything, the mass spectrometer is instructed to look only for specific precursor and fragment ions (transitions) associated with your target protein list. By focusing instrument time on a pre-defined set of peptides, the signal-to-noise ratio is exponentially increased, allowing for the detection of trace-level proteins that are otherwise lost in the global background.
Whether you choose our high-throughput prm proteomics service or the gold-standard mrm proteomics service, we help researchers replace outdated Western Blots and ELISAs with a faster, more highly multiplexed, and rigorously quantitative antibody-free targeted proteomics service. This transition is particularly critical when working with low-abundance proteins where antibody cross-reactivity or limited binding affinity often yields ambiguous results.
Content Guide
- Why Targeted Proteomics?
- Targeted MS vs. Immunoassays
- Relative vs. Absolute Quant
- Research Challenges Solved
- Comprehensive Applications
- Service Advantages
- Technology Selection Guide
- Performance Specifications
- Assay Development Workflow
- Sample Requirements
- Quantitative Deliverables
Targeted MS vs. Traditional Antibody Methods
Standard immunoassays rely on the biological recognition of an epitope, which can be masked by protein folding or compromised by off-target binding in complex matrices like serum or plasma. Targeted mass spectrometry, conversely, relies on the physical coordinates of a peptide—its mass, charge, and fragmentation pattern. This provides a level of sequence-specific evidence that antibodies simply cannot match.
| Feature | Targeted MS (PRM/MRM) | Traditional Immunoassays (ELISA/WB) |
|---|---|---|
| Multiplexing Capacity | High (Tens to hundreds of proteins simultaneously) | Low (Usually 1 target per assay, rarely up to 10) |
| Antibody Reliance | Zero (Completely antibody-free, relies on physical mass) | Absolute (Heavily dependent on antibody specificity) |
| Specificity | Extreme (Measures exact amino acid sequences/PTMs) | Variable (Prone to cross-reactivity and off-target binding) |
| Absolute Quantification | Yes (Using internal heavy AQUA peptide standards) | Yes/Semi (Requires external recombinant protein curves) |
| Linear Dynamic Range | Broad (4 to 5 orders of magnitude) | Narrow (1 to 2 orders of magnitude) |
| Assay Development Time | Weeks (Fast synthesis of target peptides) | Months to Years (Generating and validating new antibodies) |
Relative vs. Absolute Quantification: Which One Do You Actually Need?
One of the most critical decisions in a Multiplexed Biomarker Validation Proteomics project is choosing the quantitative depth. While relative quantification is perfect for confirming trends, absolute quantification provides the definitive numerical evidence required for concentration-level reporting.
Selection Logic:
- Choose Relative Quantification when your goal is to confirm whether discovery-phase hits maintain their up/down regulation trends in a larger cohort. This is cost-effective and provides statistically significant fold-change data.
- Choose Absolute Quantification (AQUA) when you need to know the exact molar concentration (e.g., fmol/μL) of a target. This is the only viable path for PK/PD modeling, bioprocess quality assessment, and multi-site method transfers.
| Dimension | Relative Quantification | Absolute Quantification (AQUA) |
|---|---|---|
| Reference Standard | Shared pool or control sample. | Spiked-in heavy isotope AQUA peptides. |
| Data Output | Peak area ratios (Fold-change). | Exact concentrations (fmol/μL or ng/mg). |
| Inter-Lab Consistency | Low (Method/Instrument dependent). | High (AQUA standards normalize variations). |
| Ideal Project Stage | Candidate ranking & hit verification. | PK/PD modeling & Process monitoring. |
Supported by our specialized absolute quantification (aqua) service workflow, we provide high-purity heavy standards to ensure your concentration-level data is defensible and interpretation-ready.
What Research Challenges Does Our Targeted Proteomics Service Solve?
From resolving positional isomers to bypassing antibody bottlenecks, see how targeted MS addresses validation roadblocks.

The "Missing Antibody" Roadblock
Bypass antibodies entirely by measuring exact peptide masses, providing definitive proof for novel variants or mutation sites.

High Background in Complex Matrices
Using ultra-fast quadrupole isolation and ion mobility, we filter out chemical noise to ensure clean signal extraction for trace analytes.

Batch-to-Batch Cohort Variability
AQUA internal standards spiked into every sample normalize run-to-run variations, maintaining statistical power across large translational cohorts.
Redundant Assay Costs
Multiplex up to 150 targets in a single run, drastically reducing cost-per-data-point and conserving precious sample volumes compared to individual ELISA kits.
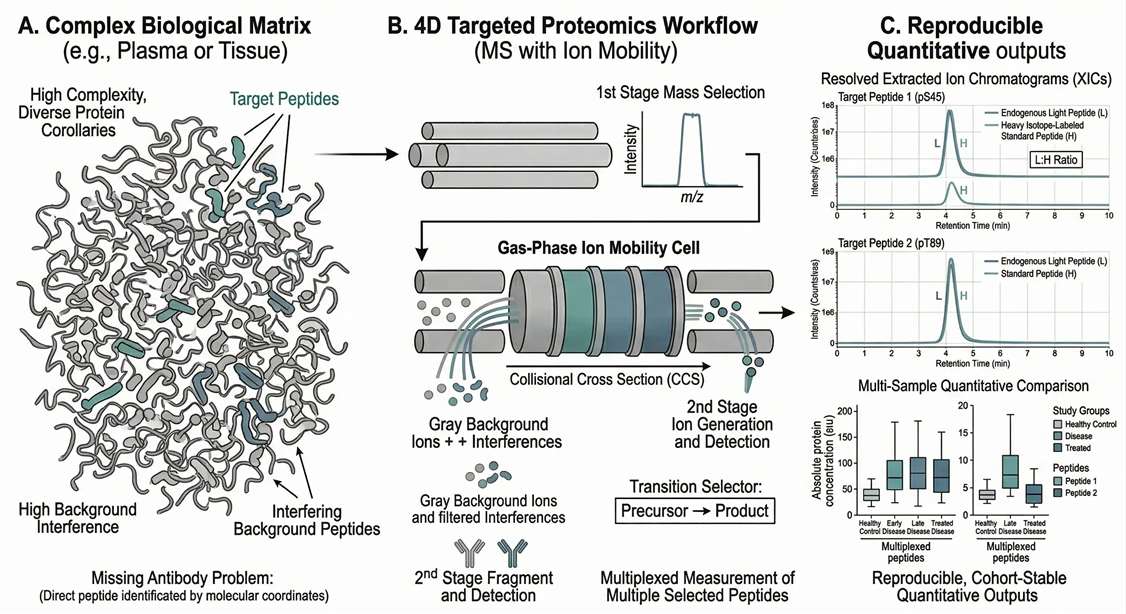
Comprehensive Applications of Targeted Proteomics
Biomarker Validation
Transition seamlessly from global discovery to establish robust candidate biomarker panels across large independent cohorts. We help translate thousands of DIA features into a manageable, validated panel for translational study packages.
Pharmacokinetics & Metabolic Pathways
Measure protein therapeutics or monitor dynamic expression via targeted metabolic pathway analysis. This is essential for monitoring enzyme induction or inhibition during lead optimization.
Bioprocessing & CMC Quality Control
For recombinant protein or viral vector production, we develop highly sensitive targeted panels to quantify residual Host Cell Proteins (HCPs) down to ppm levels. Targeted MS can detect specific deleterious HCPs that are "blind" to generic ELISA kits, ensuring rigorous process monitoring and quality assessment.
Targeted PTM Analysis
Confidently validate the stoichiometry and occupancy of specific modifications through protein post-translational modification analysis. We can design transitions that specifically target a phosphorylation, acetylation, or ubiquitination site, providing clear evidence of signaling pathway activation.
Advantages of Our Multiplexed Biomarker Validation Proteomics
End-to-End Assay Design
You provide the protein list; we handle the rest—from in silico peptide selection to standard synthesis and transition optimization.
Antibody-Free Independence
Eliminate antibody generation costs. We directly quantify sequence-specific peptides, easily distinguishing highly homologous isoforms.
Unmatched Multiplexing
Maximize precious samples. Confidently multiplex 50 to 150 distinct protein targets in a single LC-MS/MS injection.
Exceptional Sensitivity
Detect trace biological regulators with limits of detection (LOD) extending down to the attomole range, even in complex matrices.
Absolute AQUA Accuracy
Spiked heavy-isotope standards deliver true molar concentrations for accurate PK/PD modeling and technical reporting.
Rigorous Data Packages
Stringent QC protocols ensure your data is robust and ready for translational study packages and high-impact publication support.
Technology Selection Guide: MRM vs. PRM vs. Targeted 4D-PRM
Not all validation projects are identical. As an elite Targeted 4D-PRM Proteomics CRO, we match your specific panel size and matrix complexity to the optimal mass spectrometry platform.
| Dimension | MRM / SRM (Triple Quadrupole) | Standard PRM (Orbitrap / Q-TOF) | 4D-PRM (prm-PASEF via timsTOF) |
|---|---|---|---|
| Mass Analyzer | QTRAP 6500+ | Orbitrap Exploris 480 / Q Exactive | timsTOF Pro 2 |
| Multiplexing Limit | ~50–100 targets per run | ~50–150 targets per run | >200 targets per run |
| Sensitivity Focus | Extreme sensitivity for small panels | High resolution for moderate background | Maximum selectivity using Ion Mobility (CCS) |
| Assay Setup Speed | Slower (Requires extensive tuning) | Fast (Tuned post-acquisition) | Fast (Captures all fragments + mobility) |
| Best Used For | Gold-standard absolute quantitation. | Validating discovery hits; biomarker panels. | Extremely complex matrices (plasma) and PTM isomers. |
Method selection should be driven by panel size, matrix complexity, quantitative objective, and whether the project is in candidate verification, cohort expansion, or concentration-level reporting.
Selection Strategy:
- Choose MRM/SRM when the panel is tightly defined (e.g., <20 proteins) and extreme sensitivity/reproducibility is the priority.
- Choose standard PRM when panel flexibility and easier assay expansion matter more.
- Choose 4D-PRM (prm-PASEF) when dealing with extremely complex matrices, severe plasma interference, or when co-eluting background makes additional ion mobility selectivity valuable.
Assay Performance & Validation Specifications
By leveraging the advanced platforms above, we hold our targeted assays to strict analytical chemistry standards to ensure your validation data is unassailable:
- Linear Dynamic Range (LDR): ~4 to 5 orders of magnitude.
- Reproducibility (CV): Technical replicates consistently show CVs < 10%; Biological cohort CVs typically < 15%.
- Standard Curve Linearity: R² ≥ 0.99 for all synthesized AQUA standards across the reported range.
- Sensitivity: Low attomole to femtomole limit of quantification (LOQ) depending on the specific peptide target and matrix complexity.
End-to-End Custom PRM/MRM Assay Development Service Workflow
Developing a targeted MS assay is a complex process. We remove the burden with a fully managed Custom PRM/MRM Assay Development Service.
We select "proteotypic" peptides unique to your target and fly well in the MS. We screen for common PTMs or SNPs that might interfere with quantification.
We synthesize heavy-isotope labeled (AQUA) peptides and empirically determine the best transitions (precursor-to-product ion pairs) and collision energies.
We spike standards into your actual matrix to generate a multi-point calibration curve (R² > 0.99) and determine the LOD/LOQ.
Endogenous peptide signals are normalized against standards to calculate precise absolute concentrations, followed by rigorous statistical analysis.
Sample Requirements for Target Verification
Targeted proteomics is exceptionally resilient to matrix effects. In many cases, high-abundance protein depletion (which can cause variable sample loss) is not required.
| Sample Type | Recommended Input | Preservation State | Recommended Use Case | Recommended Mode |
|---|---|---|---|---|
| Fresh Frozen Tissue | 2-5 mg | Flash-frozen immediately | Tissue biomarker profiling | PRM / MRM |
| FFPE Tissue | 5-10 curls (10μm) | Mounted or Unmounted | Translational panel validation | PRM / 4D-PRM |
| Cells / Organoids | 106 cells | Washed cell pellet | Mechanistic signaling studies | PRM / MRM |
| Serum / Plasma | 10-50 μL | Unhemolyzed, spun | Biofluid biomarker verification | PRM / 4D-PRM |
| Purified Protein / HCP | 10-50 μg | Cold or frozen | Bioprocess / CMC QC | MRM / PRM |
Actionable Insights: Quantitative Deliverables and Assay Reports
Our reporting workflow focuses on analytical performance, quantitative clarity, and validation-ready interpretation rather than discovery-style feature exploration.
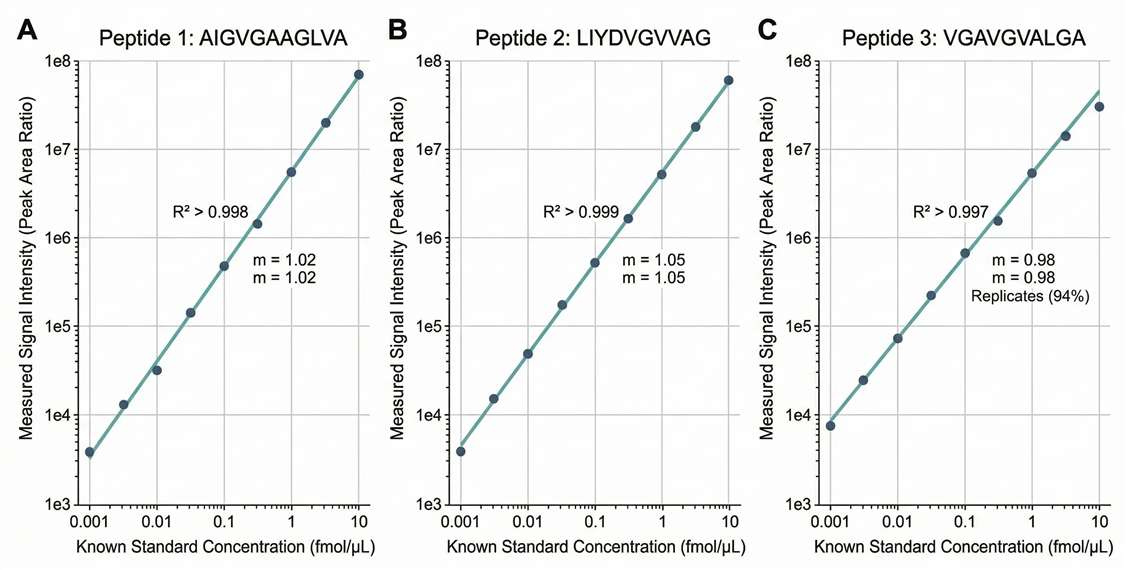
Calibration Curves: Rigorous linear regression plots demonstrating R² > 0.99 for your target peptides across the dynamic range.
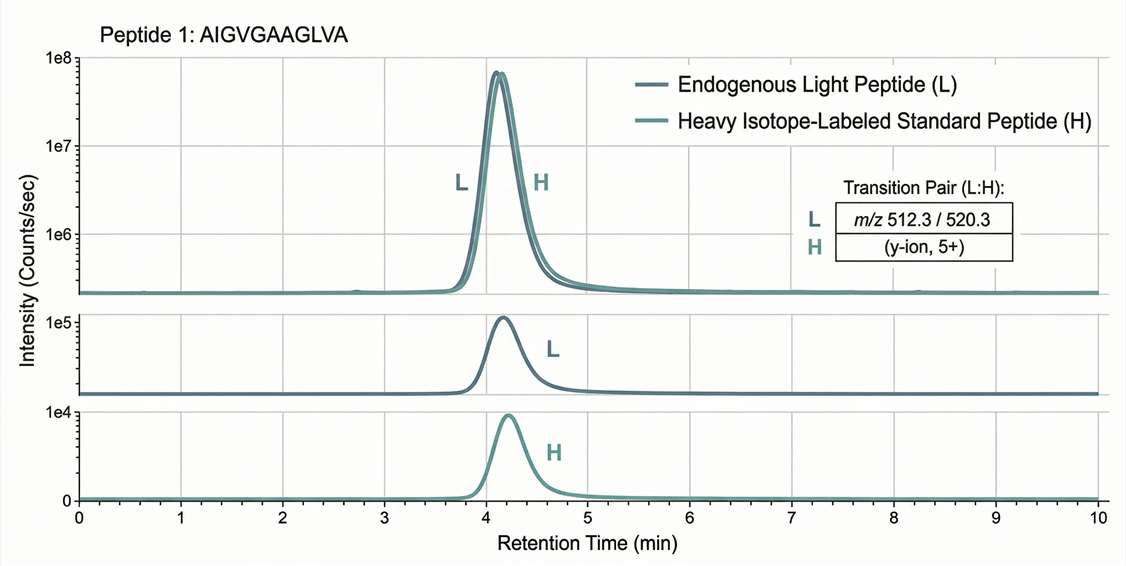
Extracted Ion Chromatograms (XIC): Clear visual evidence of co-eluting endogenous (light) and standard (heavy) peaks, confirming peptide identity.
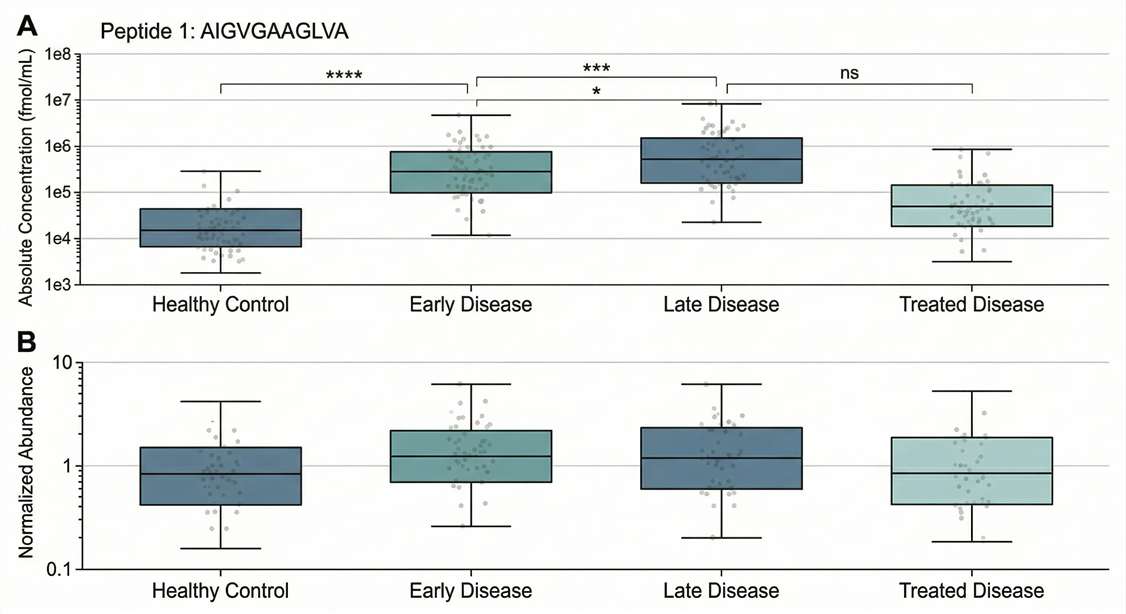
Cohort-Level Quantification: Box plots visualizing absolute concentration differences between healthy and diseased groups.
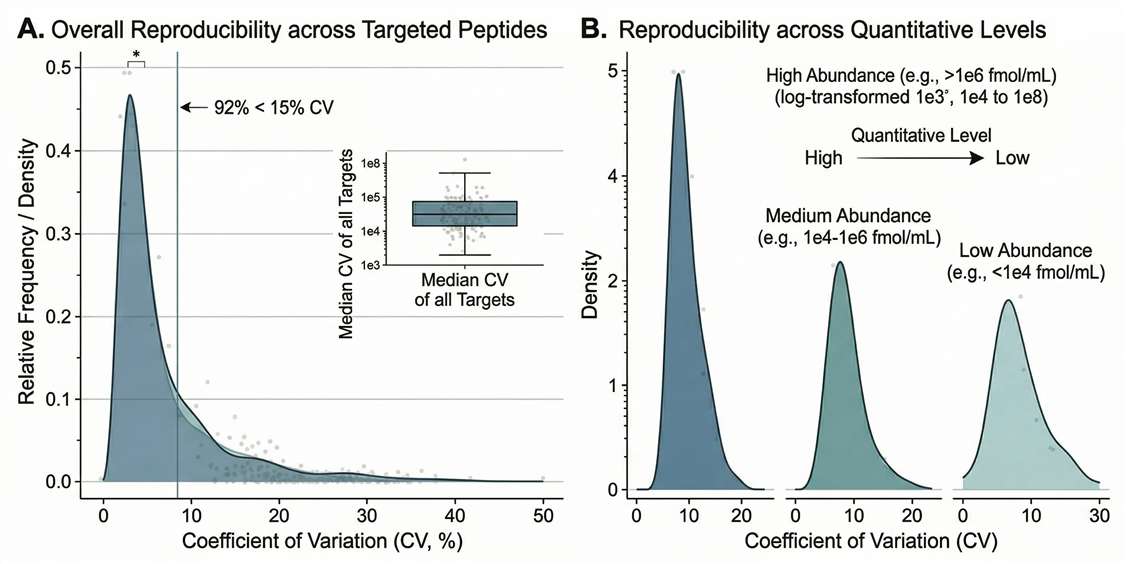
Reproducibility Metrics: Bar charts or density plots confirming technical CVs remain tightly controlled across the entire study.
Raw Data Files
Native instrument files (.d, .raw) for your own archival and secondary verification.
Processed Quantitative Tables
Excel/CSV matrices containing calculated absolute concentrations (fmol/μL or ng/mg) for all samples.
Assay Methodology
Full documentation of optimized retention times, collision energies, and precursor/fragment transitions.
Quality Control Pack
Detailed reports on assay performance, including LOD, LOQ, CV distributions, and curve linearity metrics.
Technical Reports
Statistical significance testing, group comparisons, and publication-ready figures for your targeted panels.
Antibody-Free Targeted Proteomics for Sensitive Protein Quantification
Parallel reaction monitoring targeted mass spectrometry as a fast and sensitive alternative to antibody-based protein detection
Journal: Frontiers in Analytical Science · Published: 2024
Study Scope
Traditional antibody-based methods such as Western blotting and ELISA often face limitations in sensitivity, quantitative precision, and target specificity, especially when low-abundance proteins or complex biological matrices are involved. In this study, researchers evaluated whether Parallel Reaction Monitoring (PRM) mass spectrometry could serve as a faster, more sensitive, and more quantitative alternative for targeted protein detection.
- The team developed a PRM workflow on an Orbitrap platform for targeted peptide measurement.
- They benchmarked PRM directly against immunoblotting using a recombinant target protein.
- The study assessed detection sensitivity, quantitative behavior, and the effect of complex biological background on targeted measurements.
- Heavy isotope-labeled AQUA peptides were incorporated to support more accurate relative and absolute quantification.
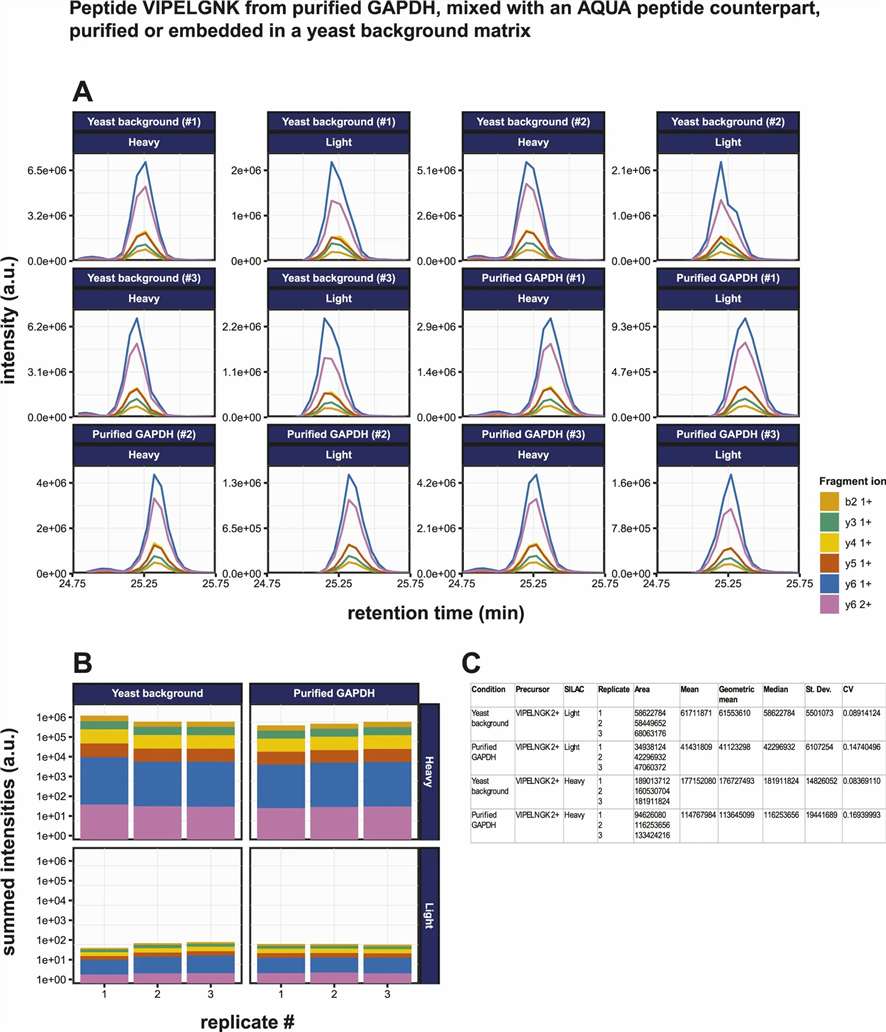 Figure 5. PRM demonstrates strong sensitivity and quantitative performance for targeted peptide detection, including in a complex background matrix.
Figure 5. PRM demonstrates strong sensitivity and quantitative performance for targeted peptide detection, including in a complex background matrix.
Quantitative Performance and Absolute Measurement
The results showed that PRM offered clear analytical advantages over traditional antibody-based detection for targeted protein measurement. The study demonstrated that PRM could detect proteolytic peptides of the purified target protein in the mid- to low-attomole range, while still maintaining useful sensitivity when the target was embedded in a complex biological matrix.
- Higher analytical sensitivity: PRM outperformed immunoblotting in low-level target detection by nearly an order of magnitude in several test cases.
- Robust quantitative behavior: Regression analysis supported stable and interpretable signal response across a 4-log concentration range, providing far superior linearity compared to the non-linear saturation common in densitometry.
- Improved specificity: PRM measured exact peptide masses and fragment ions, avoiding the "ghost bands" and cross-reactive background issues common in antibody-based methods.
- AQUA-enabled quantification: The use of heavy isotope-labeled peptides enabled more accurate relative and absolute quantification of the target analytes, providing defensible concentration values.
These findings are especially relevant for targeted proteomics projects where sensitivity, sequence-level specificity, and concentration-based reporting matter more than broad protein discovery.
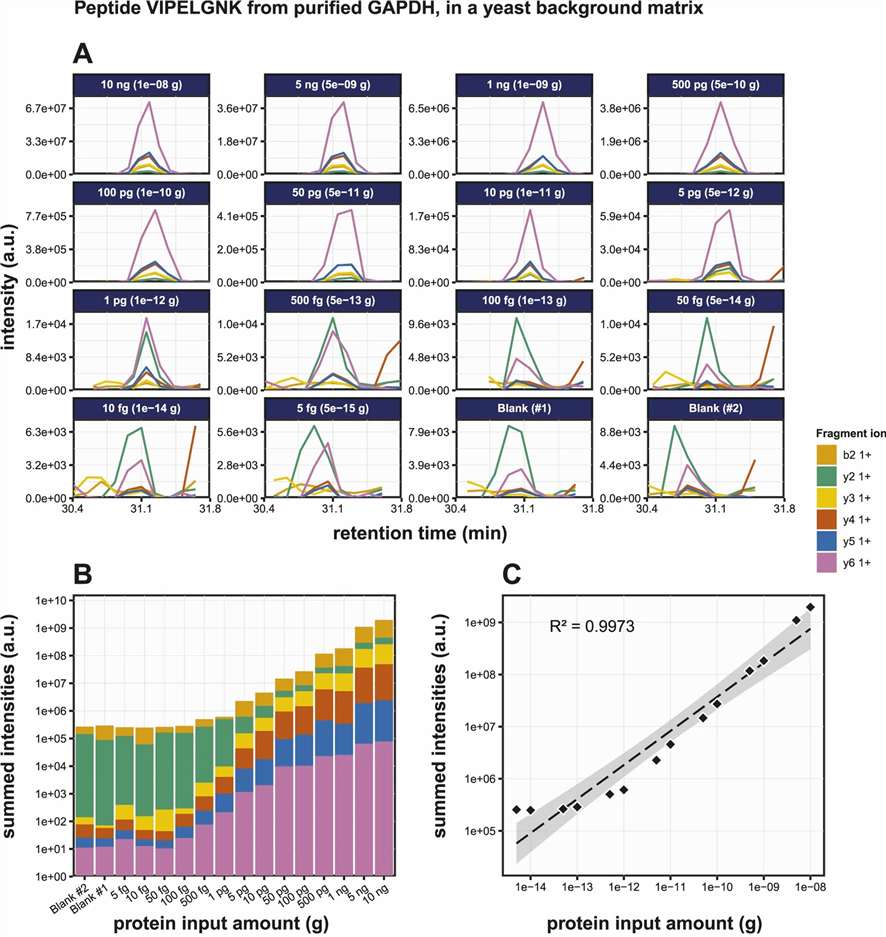 Figure 6. AQUA peptide spiking supports accurate relative and absolute quantification, with repeatable targeted measurement across replicate analyses.
Figure 6. AQUA peptide spiking supports accurate relative and absolute quantification, with repeatable targeted measurement across replicate analyses.
Why This Case Matters for Your Validation Strategy
This study provides proof-of-principle evidence that PRM can serve as a sensitive and quantitative antibody-free alternative for targeted protein measurement. For translational teams facing antibody bottlenecks, low-abundance targets, or the need for concentration-level reporting, it shows why targeted mass spectrometry can be a strong fit for validation-stage workflows.
- Supports antibody-free verification of predefined protein targets.
- Improves sensitivity for low-abundance targeted measurements in complex backgrounds.
- Enables sequence-specific quantification with reduced background ambiguity.
- Demonstrates how AQUA standards strengthen relative and absolute quantification workflows for publication-ready data.
Reference
Bezstarosti, K., Van der Wal, L., Doff, W. A. S., and Demmers, J. A. A. "Parallel reaction monitoring targeted mass spectrometry as a fast and sensitive alternative to antibody-based protein detection." Frontiers in Analytical Science 4 (2024).

