Proteomics Solutions of Drug Mechanism of Action
Small molecule drugs obtained from synthetic compounds or natural products exhibit therapeutic effects by specifically binding to one or several target proteins and regulating their functions. Conversely, bad drug-protein interactions may cause harmful side effects. Therefore, a systematic understanding of the mechanism of action of drugs at the molecular level is essential for successful clinical development. Comparing the difference in proteomic expression between cells or tissues in health and disease states makes it possible to identify the mechanism of action of small molecule drugs and characterize molecular pathways. Analyzing how disease-related proteins aggregate, identification of the interaction partners of native and aggregated pathological proteins, and determination of the consequences of protein aggregation will likely elucidate the patho-mechanisms.
Using Proteomics to Reveal the Mechanism of Drug Action
Proteomics is a large-scale study of proteins, with special emphasis on the structure and function of proteins. Proteins are an important part of organisms. They are the main components of cell metabolism and signaling pathways, and they play a very important structural role at the same time. Mass spectrometry technology has been widely used in proteomics research due to its advantages of high sensitivity, high throughput, and good compatibility.
Quantitative proteomics enables complex workflows to study the mechanisms by which small molecule drugs interact with the proteome and to measure changes in protein thermal stability following drug treatment, allowing for an understanding of direct targets and downstream regulatory events. These methods also provide powerful tools for exploring small molecule second messengers, signaling lipids, and other signaling pathways that regulate metabolites.
Our Mechanism of Drug Action Research Service
Our mechanism of drug action research service provides a broader and deeper understanding of small molecule drug targets and associated signaling pathways. The service identifies specific drug-protein interactions, as well as the pharmacological effects produced by drug candidates. The application is based on our proprietary Discovery Proteomics - high-throughput, label-free quantitative DIA proteomics technologies.
Workflow
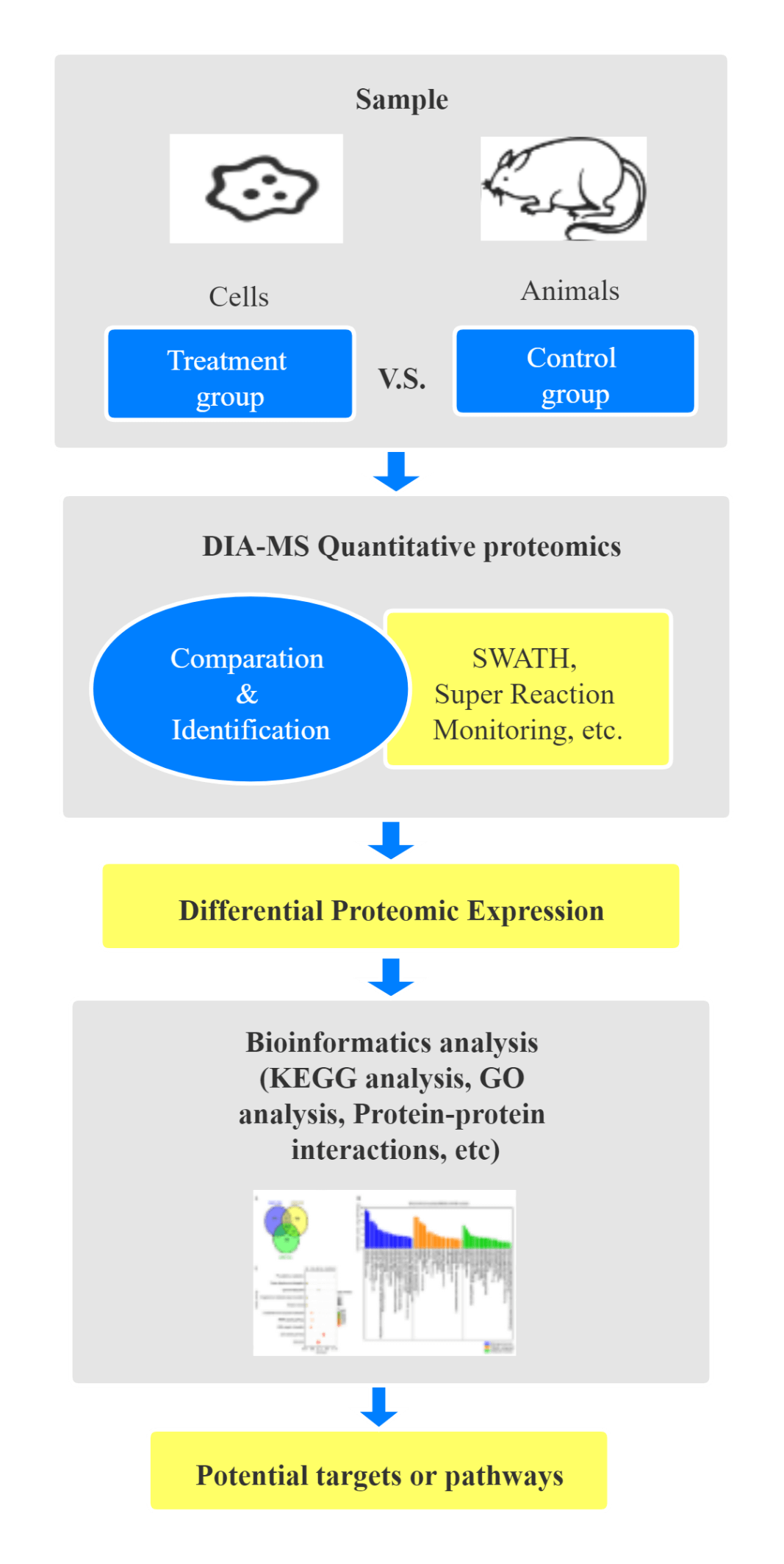 Schematic diagram of the experimental procedure for differential proteomics in studying of the mechanism of drug action
Schematic diagram of the experimental procedure for differential proteomics in studying of the mechanism of drug action
Analytical Platforms
AB SCIEX Triple-TOF 5600-plus, Q-Exactive, Orbitrap Fusion
Key Features
- High accuracy
- High throughput, more than 9000 proteins can be identified and quantitated at once
- Quantitatively identify nearly all detectable molecules, covering low abundance proteins/peptides
- High repetition rate
- Complete and comprehensive information storage of samples in the first analysis
- High repetition rate
Sample Requirements
Our special protein extraction technology allows us to quickly extract proteins from various samples and design personalized experimental schemes according to different experimental purposes. Specific requirements are as follows:
| Sample Type | Protein | # of Cell |
| Quantify | 100 ug | 1×107 cells |
Note
Please prepare enough dry ice or ice packs to ensure low temperature during sample.
Delivery
Our service is for research use only and is not intended for diagnosis.
Report
- Experimental steps
- Relevant parameters
- Mass spectrometry spectra
- Raw data
- Proteomics analysis results
References:
- Thomas J. Hedl et al. Proteomics Approaches for Biomarker and Drug Target Discovery in ALS and FTD. Front Neurosci. 2019; 13: 548.
- Tao Cui et al. Uncovering Drug Mechanism of Action by Proteome Wide- Identification of Drug-Binding Proteins. Med Chem. 2017;13(6):526-535.
- Shuailong Jia et al. Elucidation of the Mechanism of Action for Metal Based Anticancer Drugs by Mass Spectrometry-Based Quantitative Proteomics. Molecules. 2019; 24(3): 581.

 4D Proteomics with Data-Independent Acquisition (DIA)
4D Proteomics with Data-Independent Acquisition (DIA)