The endoplasmic reticulum (ER), as a unique multifunctional organelle, is involved in important physiological processes such as lipid biosynthesis, protein folding and modification, Ca2+ dynamic maintenance and cellular detoxification. The communication between ER and other organelles (e.g. mitochondria, Golgi apparatus, etc.) plays a key role in a variety of cellular functions.
Endoplasmic reticulum dysfunction caused by genetic or environmental damage leads to endoplasmic reticulum stress, which activates the unfolded protein response. The unfolded protein response is a cellular self-protective measure, but excessive or prolonged endoplasmic reticulum stress can cause apoptosis. Therefore, the development of some diseases is closely related to the physiological abnormalities of the endoplasmic reticulum. Studies have demonstrated that endoplasmic reticulum stress signaling is involved in cancer, inflammatory diseases, metabolic diseases, osteoporosis and neurodegenerative diseases. The comprehensive identification and characterization of ER-related proteins and metabolites are important not only for basic biological research but also for studying and revealing the etiology of diseases.
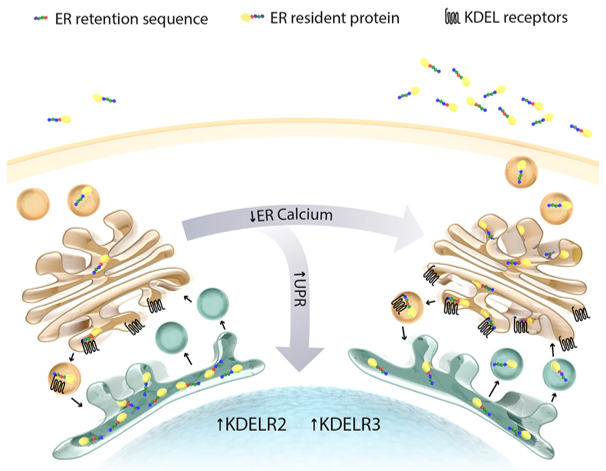 ER and KDEL receptors (Trychta et al., 2018)
ER and KDEL receptors (Trychta et al., 2018)
In plant studies, misfolded and unfolded proteins accumulate in large quantities in the endoplasmic reticulum when plants are under adversity, leading to endoplasmic reticulum stress. To alleviate stress, cells initiate conserved unfolded protein response pathways to help protein folding or degrade misfolded proteins. Studying endoplasmic reticulum stress response can help in the exploration of plant development, stress resistance, disease resistance and other processes.
Creative Proteomics provides you with a professional endoplasmic reticulum multi-omics analysis platform. We have high-resolution mass spectrometry instruments and a professional bioinformatics team to provide you with endoplasmic reticulum proteomics, endoplasmic reticulum metabolomics and endoplasmic reticulum lipidomic analysis services to promote the functional mechanism research of endoplasmic reticulum in various fields of human, animal and plant, etc. We aim to provide you with high-quality services to accelerate the progress of your project and open up new horizons.
The Endoplasmic Reticulum Assays We Offer Include but Are Not Limited to:
1. Endoplasmic reticulum isolation and purification service
2. Endoplasmic reticulum proteomics analysis
- Study of protein expression patterns: study of the characterization of all proteins in a particular cell or tissue under specific conditions
- Study of protein function patterns: reveal all protein functions and their modes of action
- Study of protein-protein and protein-DNA interactions
- Analysis of post-translational modifications of Endoplasmic reticulum proteins
- Structural analysis of proteins
- Subcellular localization of proteins
3. Endoplasmic reticulum metabolomics analysis
- Endoplasmic reticulum metabolite differential expression analysis
- Qualitative and quantitative analysis of endoplasmic reticulum metabolites
4. Endoplasmic reticulum lipidomic analysis
- Analysis of Endoplasmic reticulum lipid composition and levels
- Analysis of differential expression of Endoplasmic reticulum lipid molecules
- Qualitative and quantitative analysis of Endoplasmic reticulum lipid molecules
- Endoplasmic reticulum membrane fatty acid analysis
Basic Analysis
- Data quality control and molecular identification
- Expression quantitative analysis
- Differential analysis
- Functional annotation and enrichment analysis
Reference
- Trychta, K. A., Bäck, S., Henderson, M. J., & Harvey, B. K. (2018). KDEL receptors are differentially regulated to maintain the ER proteome under calcium deficiency. Cell reports, 25(7), 1829-1840.
* For Research Use Only. Not for use in diagnostic procedures.