After transcription and translation, genes produce long chains of amino acids with specific sequences, which are precursors to proteins. After covalent modification, folding, curling, and forming a specific spatial conformation, they become mature and functional proteins. Covalent modification plays an important role in the protein maturation process. Many key functions of proteins, especially those related to regulation, are achieved after covalent modification. Post-translational modifications can occur at any site of a protein. There are currently hundreds of post-translational modifications reported in the literature. Common ones include glycosylation modification and acetylation modification.
The Role of Glycosylation Modification
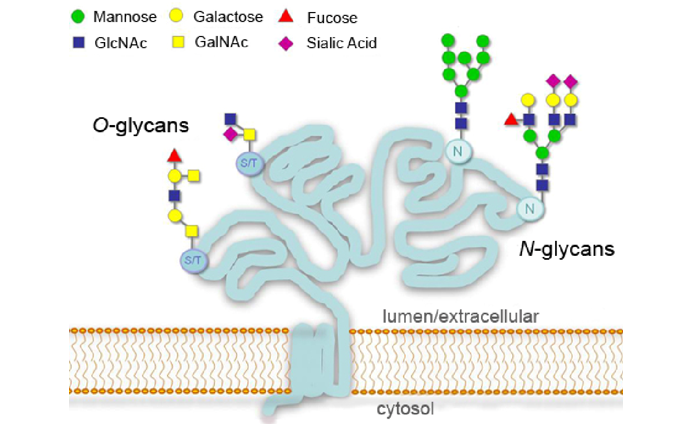
Protein glycosylation is one of the most extensive, complex, and important post-translational modifications. It is inferred that more than 50% of the proteins have undergone glycosylation modification. These glycoproteins are widely distributed in living bodies, especially abundant in cell membranes and body fluids. Most membrane proteins and secreted proteins are glycoproteins. Glycosylation modification not only affects the spatial conformation, biological activity, transport and localization of proteins, but also plays an important role in specific biological processes such as molecular recognition, cell communication, and signal transduction.
The zona pellucida on the surface of vertebrate egg cells is a specific type of glycoprotein that is essential during the process of sperm egg recognition, adhesion, and membrane fusion. The sperm receptor glycoprotein is a key molecule that induces sperm-egg binding. Only when the sperm is combined with a high degree of specificity can the acrosome reaction be induced to achieve fertilization.
In addition, most of the molecules involved in the immune response are glycoproteins, such as immunoglobulins, cytokines, the complement system, and leukocyte differentiation antigens. Their sugar chain structure is involved in B cell activation, T cell activation and apoptosis, antigen processing and presentation, and affects the structure and function of immunoglobulins. T cell also plays a role in regulating lymphocyte activation and apoptosis, antigen recognition and clearance, cell adhesion, signal transmission, and endocytosis.
Techniques used in Protein Glycosylation Research
Protein glycosylation research can be divided into three parts: glycoprotein analysis, glycosylation site analysis, and sugar chain analysis.
Traditional research methods include separation analysis by chromatography or electrophoresis, X-ray crystal diffraction or nuclear magnetic resonance (NMR) structural analysis, and functional analysis by biological methods. However, due to the high complexity of glycosylation modification, these methods are cramped and limited.
High-throughput biomass spectroscopy has been more developed and hence, became widely used in glycosylation research. Since the 1980s, the development of soft ionization technology represented by the matrix-assisted laser description ionization (MALDI) and electrospray ionization (ESI) has made the analysis on biological macromolecules with molecular weights of tens or even hundreds of thousands at pmol (10-12mol) to fmol (10-15 mol) levels more accurately. This technology brings a breakthrough for the application of mass spectrometry in the research of biological macromolecules.
- Qualitative methods of protein glycosylation
Qualitative study of protein glycosylation based on biomass spectrometry mainly includes identification of glycosylation sites and analysis of sugar chain structure. The major steps of qualitative research on protein glycosylation includes:
1. Separation and enrichment of glycopeptides.
At present, the most effective separation and enrichment techniques are the lectin method based on affinity action and the hydrazone reaction method based on covalent action.
2. Enzymatic digestion of glycopeptide sugar chain
3. Sample preparation for mass spectrometry analysis.
In addition to enrichment and enzymatic hydrolysis, sample preparation processes such as protein denaturation, desalination, and high abundance removal are also the basis and key to mass spectrometry analysis. The purpose of protein denaturation is to destroy the three-dimensional structure of the protein. The resultant stretched-out protein is beneficial for the enrichment and enzymatic hydrolysis to proceed more fully. Mass spectrometry is very sensitive to salt. The non-volatile salts commonly used in biological sample processing will inhibit the ionization of the sample, resulting in reduced detection sensitivity or even failed detection. Therefore, desalting is required before mass spectrometry. In special samples such as serum, a few proteins often account for the vast majority of the total amount, inhibiting the effective analysis of a large number of low-abundance target proteins. Therefore, it is necessary to remove highly abundant proteins.
4. Identification of glycosylation sites
5. Sugar chain structure analysis: sugar chain structure analysis techniques include NMR, chromatography, chip technology, etc. Using mass spectrometry to analyze sugar chain structure is popular because of its high sensitivity and accuracy, which is conducive to the analysis of trace and complex mixed samples.
After the sugar chain is fragmented in series (MS2) or multi-level (MSn), the glycosidic bond is broken or the ring is broken according to a certain rule. By analyzing the fragment ions, the composition and structure of the sugar chain can be inferred, especially the fine structures such as branching, glycosidic bonds, and conformation.
- Quantitative methods for protein glycosylation
Quantitative techniques for protein glycosylation are based on qualitative research techniques. By comparing the peak intensity and peak area of parallel samples, the change characteristics of the quantity are analyzed. Quantitative methods based on mass spectrometry can be divided into semi-quantitative, relative quantitative and absolute quantitative. Semi-quantitative means adding a standard as an internal standard, or using analysis software and other means to compare peaks with specific mass-to-charge ratios in different spectra. Relative quantification, by isotopically labeling peptides, sugar chains and other analytes, causes the difference in the quality on analytes of different samples, so that different samples can be compared in the same spectrum. Absolute quantification uses the selective reaction monitoring (SRM) or multiple reaction monitoring (MRM) developed in recent years to determine the absolute content of specific proteins in samples.
At present, quantitative analytic methods can be divided into label-based and non-label-based. Several common isotope labeling quantitative techniques include isotope-coded affinity tags (ICAT), isobaric tag for relative and absolute quantitation (iTRAQ), and stable isotope labeling by amino acids in cell culture (SILAC). Non-labeled quantification is mainly based on mass spectrometry technology, including SRM, MRM, SWATH, etc.
References:
1. Magnelli P E, Bielik A M, Guthrie E P. Identification and characterization of protein glycosylation using specific endo-and exoglycosidases. JoVE (Journal of Visualized Experiments), 2011 (58): e3749.
2. Mariño K, Bones J, Kattla J J, et al. A systematic approach to protein glycosylation analysis: a path through the maze. Nature chemical biology, 2010, 6(10): 713-723.