Following genomics and proteomics, glycomics is becoming another new frontier and hotspot in life science research. Glycomics takes the sugar group expressed in a specific space-time or in a specific physiological and pathological state of the research object, and analyzes the composition and activity of carbohydrates from a holistic perspective. The glycoprotein sugar chain structure is one of its important research contents. The sugar chain has many branching structures and spatial conformations, thus carrying a lot of biological information and playing complex functions in the living body.
According to the different attachments with sugar chains, proteins mainly have N-linked and O-linked glycosylation forms. N-linked glycosylation is the sugar chain connected to -NH2 on Asn residue in protein Asn-XXX-Ser/Thr sequencer (XXX is an amino acid other than proline). O-linked glycosylation is when the sugar chain is connected to -OH on the Ser/Thr residue of the protein. N-linked and O-linked glycoproteins are mainly located on the cell membrane surface, and can also exist as secreted proteins. In addition, there are two forms of glycoproteins, GPI anchored protein and O-GlcNAc modified protein.
Analysis Method of Sugar Chain Structure
The sugar chain structure information includes monosaccharide composition of the sugar chain, anomeric carbon configuration, glycosidic bonds linkage, and sequence analysis of sugar residues. The structure analysis of glycoprotein sugar chain usually includes the steps of extraction and separation of glycoprotein, sugar chain release and sugar chain structure identification.
- Extraction and separation of glycoprotein
Glycoproteins are mostly membrane proteins, which are located on the cell surface and have poor water solubility. It is common to prepare the plasma membrane first, then use detergents such as CHAPS or Triton X2114 to perform phase separation, organic solvent extraction and other methods to extract glycoproteins. The obtained glycoproteins are mainly separated and purified by dialysis, ultrafiltration, electrophoresis and chromatography.
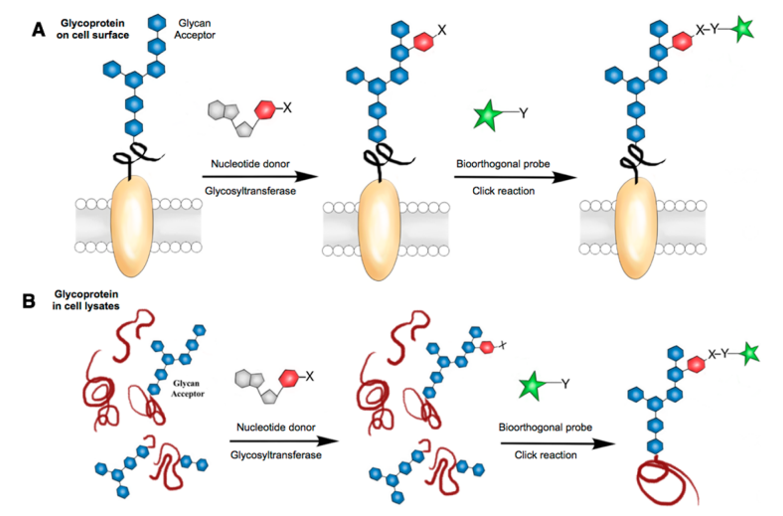
Methods are available to directly study the structure of the glycoprotein sugar chains on the surface of the entire cell without glycoprotein extraction and purification, such as Sugar Microarray and the magnetic-activated cell sorting (MACS) technology.
- The release of sugar chains
The purified glycoprotein sample is ready for sugar chain release. Sugar chain release is commonly used in chemical release and enzyme release.
The chemical methods mainly include hydrazine solution and β2 elimination.
- The hydrazine hydrolysis method can release N-sugar chains and O-sugar chains of glycoproteins using automation. However, due to the severe reaction conditions of this method, acetylation and recovery treatment of the sugar chain is required after hydrazine hydrolysis, which destroys the protein, making it impossible to study the glycosylation site.
- β2 elimination method allows the release of the complete O-sugar chain with the reducing end. At the same time, this method affects the release of N-sugar chains, which can recognize trace analysis of O-sugar chains and N-sugar chains.
The widely used enzymatic method for releasing glycoprotein sugar chains is simple and the conditions are mild. It provides information on the composition, arrangement order and α or β configuration of glycosidic residues of sugar chain residues. The more commonly used specific proteases that release N-sugar chains are N-glycopeptidase F (PNGase F) and N-glycopeptidase A (PNGase A).
- The label of the sugar chain
In order to detect sugar chains more effectively in the process of separation, purification, and structure identification, it is often necessary to label or derive sugar chains. It is necessary for the separation and analysis of trace sugar chains. Common labeling methods include radioisotope labeling, uv and fluorescence derivatization.
- Method for separating and purifying sugar chain and identifying structure
1) Chromatography
High performance liquid chromatography (HPLC), frontier affinity chromatography (FAC), high performance anion exchange chromatography (HPAEC). In order to separate and analyze the sugar chain mixture more fully and accurately, different types of HPLC can be combined into multi-dimensional HPLC (MD-HPLC), such as ion exchange HPLC, normal phase HPLC, reverse phase HPLC combination constitutes 3D-HPLC. MD-HPLC technology can provide more information due to its low peak overlap, large peak capacity, high resolution, and good selectivity. This technique is not only important for the separation and purification of glycoprotein sugar chains, but also the current mainstream technology for sugar chain structure analysis.
2) Mass spectrometry
Electrospray mass spectrometry (ESI-MS) and matrix assisted laser desorption time-of-flight mass spectrometry (MALDI-TOF-MS) were used to analyze the structure of high-polarity, volatile and thermally unstable sugars. On-line techniques, such as GC-MS, HPLC-MS, CE-MS and Chip-MS, can be used to directly analyze trace samples without too many steps of separation and purification, so as to obtain relevant sugar chain structure information more effectively.
3) Nuclear magnetic resonance (NMR)
NMR is one of the most important methods often used to analyze sugar chain stereochemical structure. It can determine the configuration, connection position, branching and micro-diversity of sugar chains.
The glycolic microarray technology represented by sugar microarray and lectin microarray can indirectly provide the structure information of sugar chain by using the specific recognition and binding ability between sugar chain and lectin. It is a good tool for high throughput and rapid analysis of protein glycation types. The prepared lectin microarray can also be further used to obtain more accurate information about sugar chain structure by biomolecular interaction analysis (BIA) and surface enhanced laser analytical ionization time-of-flight mass spectrometry (SELDI-TOF-MS).
Reference:
- Lopez Aguilar A, et al. Tools for studying glycans: recent advances in chemoenzymatic glycan labeling. ACS chemical biology, 2017, 12(3): 611-621.