iTRAQ is an acronym of Isobaric tag for relative and absolute quantitation, which was developed by Applied Biosystems Incorporation in 2004. It is an isobaric labeling method to determine the amount of proteins from different sources in just one single experiment by mass spectrometry.
The principle and workflow of iTRAQ
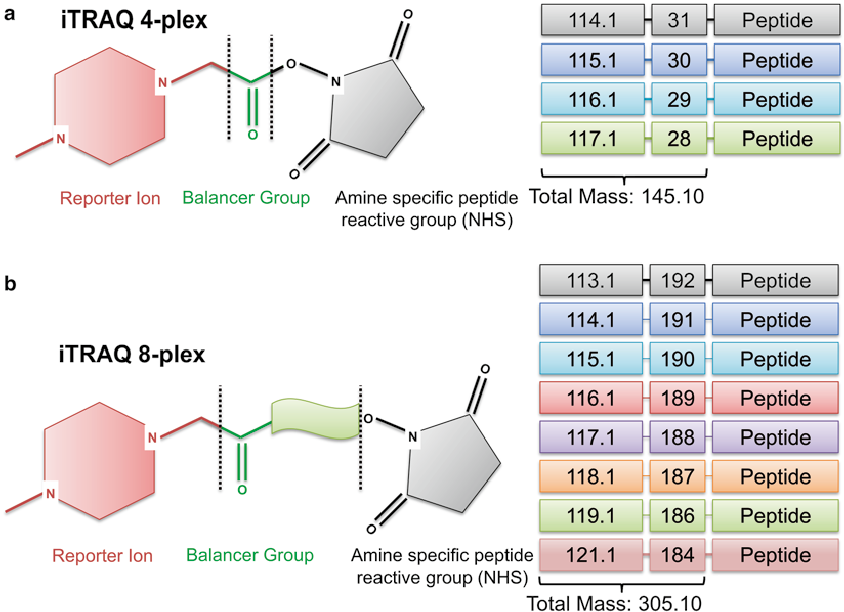
First, we should know the structure of iTRAQ reagents. The isobaric tagging reagents consist of a unique charged reporter group, a peptide reactive group, and a neutral balance group. The peptide reactive group covalently links an iTRAQ reagent isobaric tag with each lysine side chain and N-terminus group of a peptide, labeling all peptides in a given sample digest. The neutral balance group ensures the iTRAQ labeled-peptide displays the same mass to maintain an overall mass of 145 Da for 4-plex and 305 Da for 8-plex.
Figure 1. Structure of iTRAQ reagents
A mixture of peptides is obtained by hydrolyzing the protein sample. All peptides in the sample are labeled with different iTRAQ reagents, and all labeled protein samples are mixed. The peptides are subjected to tandem mass spectrometry to obtain the mass spectrum. In the first-stage mass spectrometry, after being labeled with different isotopes regardless of which iTRAQ reagent is used, the same peptide from different sources is completely identical in molecular weight and they appeared as the same peak. Samples of the same peak are collected to perform a second-stage mass spectrometry, in which the bond between the balance group and the peptide reactive group is broken and the balance group is lost. The same peptide with different isotopic labels produces different masses of reporter ions, and the reporter ions exhibit different peaks. Quantitative information of the same peptides between different samples can be obtained by analyzing related data with software and databases.
In the general workflow for an iTRAQ experiment, each sample is reduced, cysteine blocked, and digested with trypsin. Each sample is labeled with a different iTRAQ tag in a single tube. And then we can combine all iTRAQ reagent-labeled samples into one sample mixture for liquid chromatography-tandem mass spectrometry (LC-MS/MS) analysis.
Advantages and disadvantages of iTRAQ
Quantitative proteomics using mass spectrometry-based approach is one of the most important advantages of iTRAQ, especially in the field of clinical proteomics. The progress of metabolism of a particular drug or the progressive concentration of certain biomarkers at different stages of a particular disease can easily be quantified using iTRAQ-based studies. Multiplexing ability is another great advantage of iTRAQ. There are eight reporter ions available that therefore achieve multiplexing at the level of eight samples. Also, iTRAQ can reduce overall time and variation. On the other hand, iTRAQ reagents are extremely costly and also extremely sensitive to contamination from salts. Moreover, sophisticated software is required for analyzing iTRAQ data. Another disadvantage is the variability arising due to the inefficient enzymatic digestion.
An example of iTRAQ
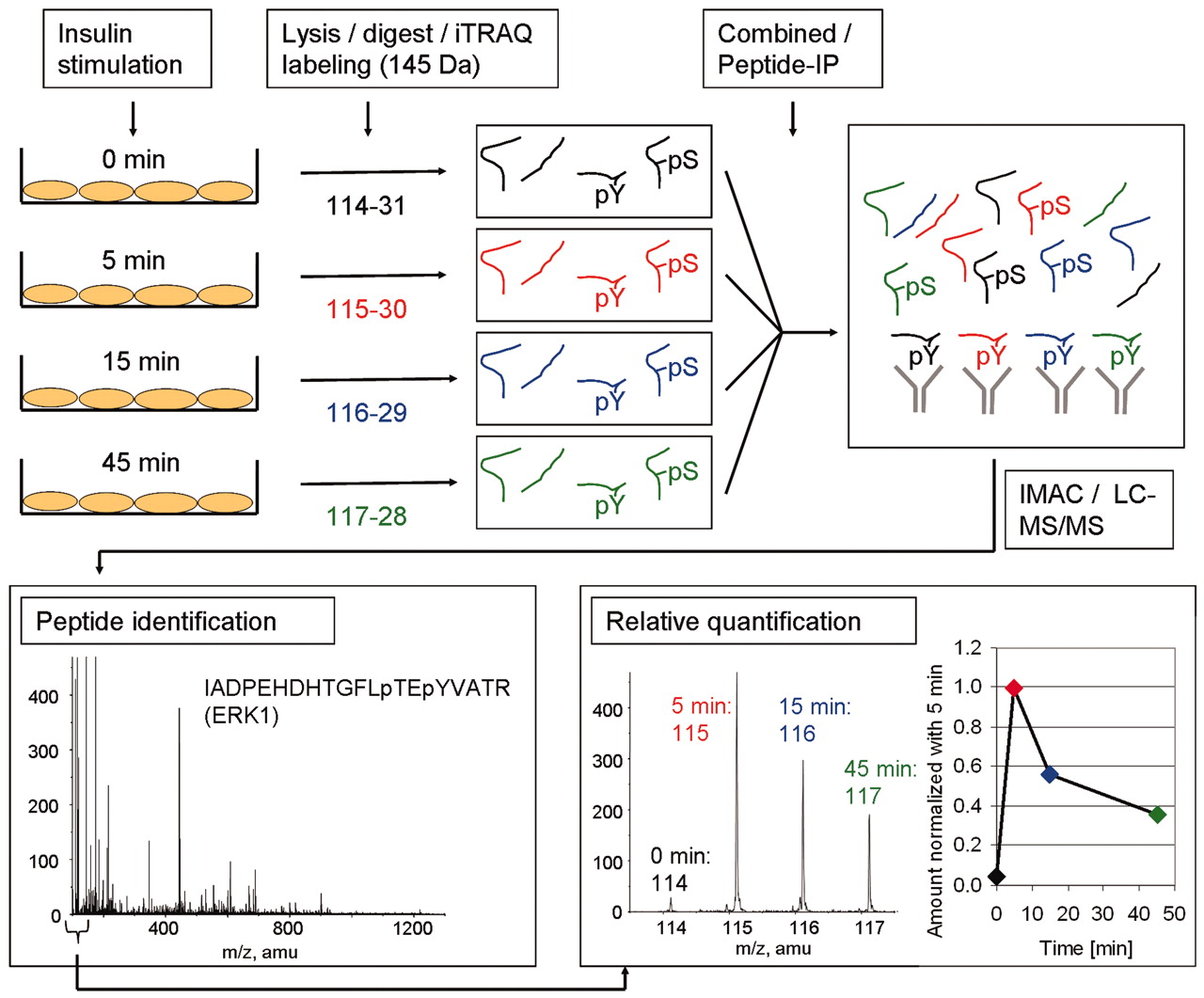
Figure 2. The Methodology used to identify and quantify tyrosine phosphorylation sites upon insulin stimulation. (Schmelzle K., et al; 2006)
There is an example that iTRAQ was used to identify and quantify tyrosine phosphorylation sites upon insulin stimulation. In this experiment, adipocytes were stimulated with insulin for different times. After cell lysis and digestion, the researchers used iTRAQ to label resulting peptides. The labeled peptides were combined for further analysis (phosphotyrosine peptide immunoprecipitation, immobilized metal affinity chromatography, and LC-MS/MS). After LC-MS/MS, the identity of the peptide was determined and the area of the tag masses 114, 115, 116, and 117 mass-to-charge ration (m/z) was used to calculate the ratio of phosphorylated peptides at the different time points compared with 5-min insulin stimulation.
iTRAQ has been widely used for quantitative analysis of proteins and in biomaterials such as microorganism, animal, plant and biomedical samples. At Creative Proteomics, we are confident to provide professional and reliable iTRAQ services and other proteomics quantification services, including but not limited to:
- iTRAQ-based proteomics analysis service
- TMT-based proteomics analysis service
- SILAC-based proteomics analysis service
- Absolute quantification (AQUA) service
- Label-free quantification service
- Semi-quantitative proteomics analysis service
Reference:
- Schmelzle K, Kane S, Gridley S, et al. Temporal dynamics of tyrosine phosphorylation in insulin signaling. Diabetes, 2006, 55(8): 2171-2179.