What is Redox Proteomics?
Redox proteomics is a specialized branch of post-translational modification (PTM) proteomics that systematically investigates redox-dependent changes in protein structure, activity, and interactions. By profiling these changes across the proteome, it helps explain how cells use redox balance to control normal functions and respond to stress.
Because redox modifications are dynamic and often reversible, their analysis relies on high-resolution mass spectrometry, selective chemical labeling, and robust data analysis. These capabilities make redox proteomics a valuable discovery tool in cell biology, pharmaceutical research, environmental studies, and bioengineering.
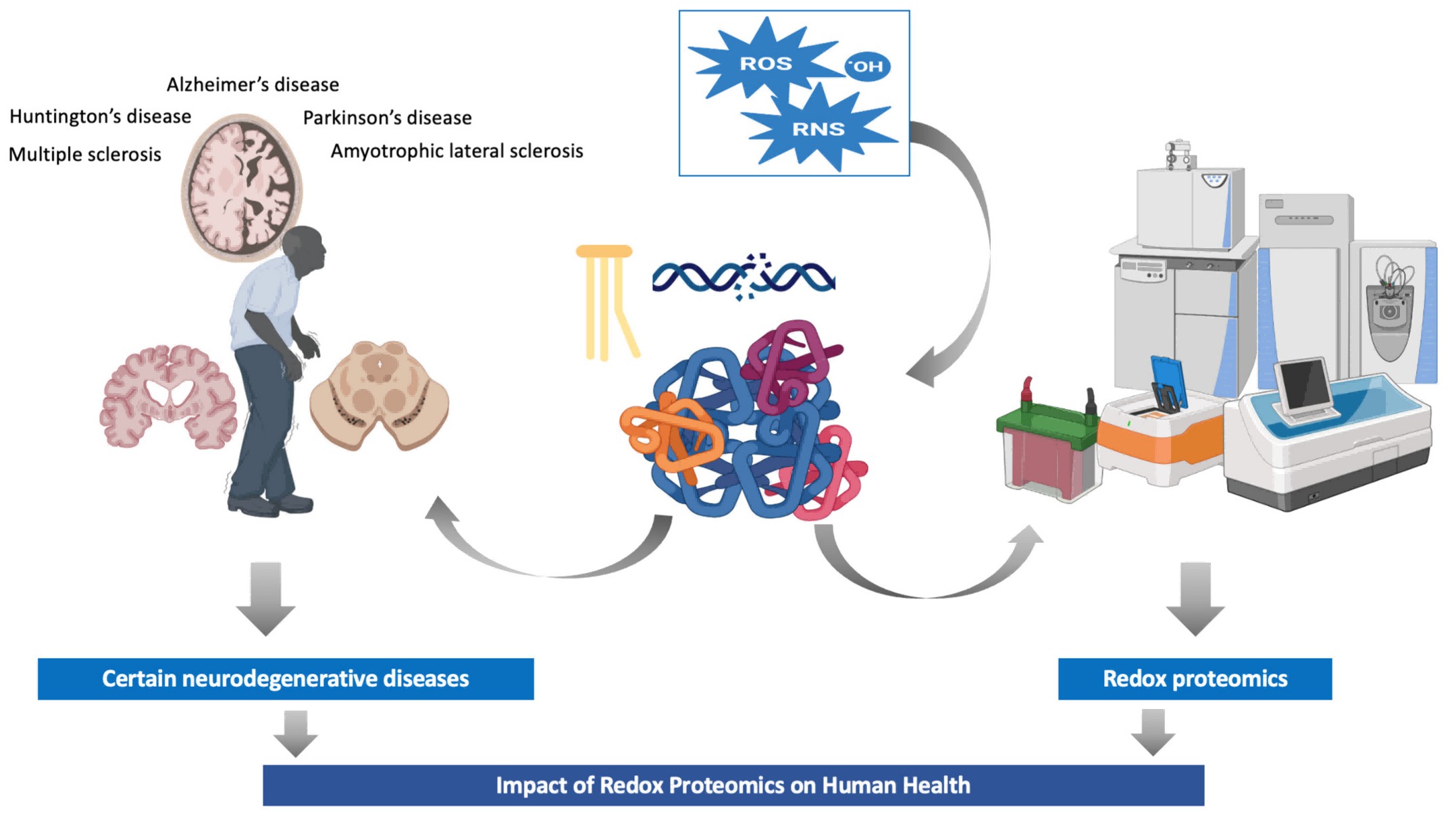
Figure 1. Redox proteomics: impact on human health and new insights (Cadenas-Garrido P, et al., 2024).
Why Redox Proteomics Requires Specialized Analytical Strategies
Conventional proteomics workflows often struggle to capture redox modifications because these modifications are chemically unstable and can be easily altered during sample handling. As a result, crucial oxidation-dependent protein features can be lost or misrepresented.
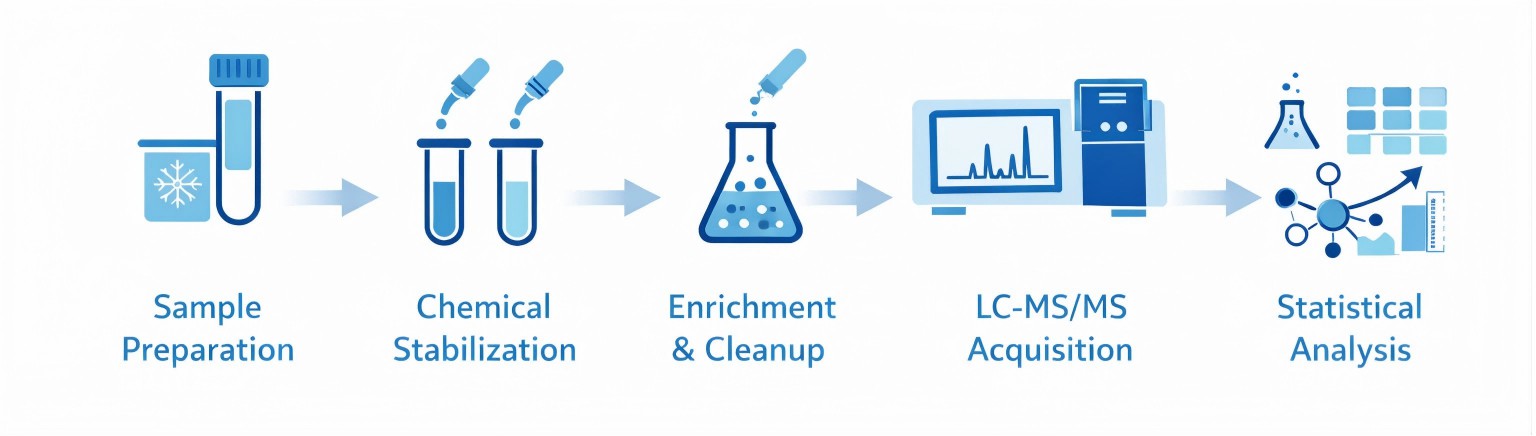
Preserving Native Redox States: Careful control of oxidation and reduction during sample handling is essential. Antioxidant-controlled preparation minimizes artificial modification while preserving endogenous redox states.
Chemical Labeling and Selective Enrichment: Differential chemical labeling, switch-based assays, and affinity capture strategies stabilize redox modifications and enrich modified peptides. These steps are critical for improving sensitivity and site-localization confidence.
PTMs Analyzed by Redox Proteomics
S-Nitrosylation
S-nitrosylation is a reversible protein modification in which nitric oxide attaches to a cysteine sulfur atom. This small chemical change can switch enzymes on or off, influence how long a protein remains stable, and fine-tune cellular signaling pathways. Using high-resolution redox proteomics, researchers can accurately pinpoint where S-nitrosylation occurs and measure how these modifications change across different biological conditions, providing clear insight into nitric oxide–driven regulation.
S-Glutathionylation
S-glutathionylation is a reversible modification in which glutathione forms a mixed disulfide bond with a protein cysteine. This process acts as a redox buffer, temporarily shielding sensitive cysteine residues from permanent oxidative damage. At the same time, it serves a regulatory role by modulating protein activity as cellular redox conditions change, thereby fine-tuning signaling pathways during oxidative stress and recovery.
Disulfide Bond Formation and Rearrangement
Both intra- and intermolecular disulfide bonds influence protein folding, complex assembly, and structural stability. Redox proteomics captures dynamic disulfide remodeling that cannot be resolved by standard proteomics alone.