What Is Membrane Proteomics and Why Does It Matter?
Membrane proteomics is a sub-discipline of proteomics dedicated to the large-scale analysis of proteins associated with biological membranes, including plasma membranes and intracellular organellar membranes. Unlike soluble proteins, membrane proteins are embedded within or tightly associated with lipid bilayers, requiring tailored enrichment, solubilization, and analytical strategies.
Understanding the membrane proteome is essential for elucidating cellular signaling networks, transport mechanisms, and protein interaction landscapes. Comprehensive membrane protein profiling enables researchers to investigate how membrane composition varies across different cell types, experimental conditions, or biological states, providing valuable insights into cellular organisation and regulation.
Why Focus on Membrane proteins?
Membrane proteins perform diverse and essential biological functions. They regulate the movement of ions and molecules across membranes, mediate signal transmission between cells and their environment, and facilitate cell–cell recognition and adhesion. These functions position membrane proteins at the interface between intracellular processes and extracellular cues.
From a research perspective, membrane proteins are of particular interest because alterations in their expression, localization, or modification often reflect changes in cellular state. Studying membrane protein dynamics contributes to a deeper understanding of biological systems at both molecular and systems levels, supporting the generation of hypotheses and the exploration of mechanisms.
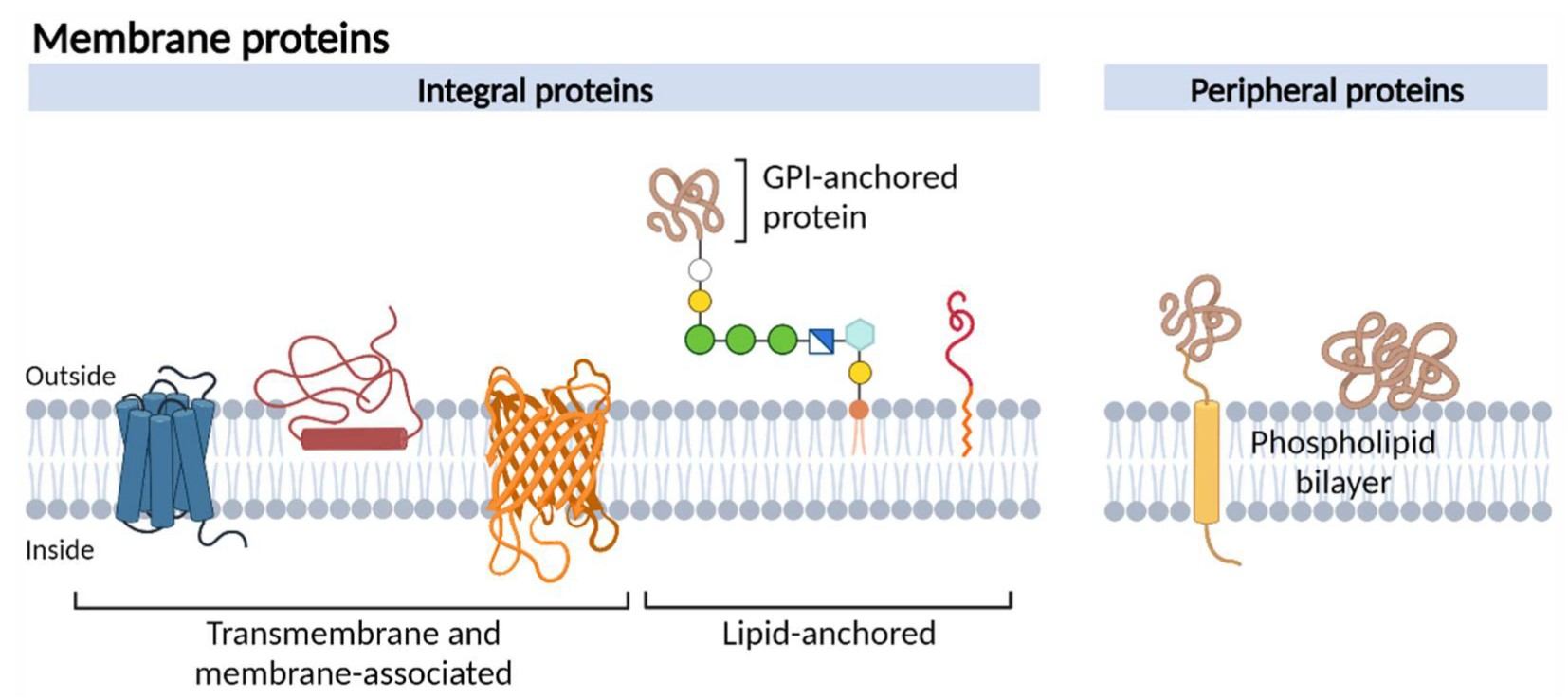
Figure 1. Schematic representation of different membrane protein types (Jong E, Kocer A. 2023).
Membrane enrichment and protein extraction strategies
Effective enrichment reduces dynamic range and improves detection of membrane proteins:
- Density gradient centrifugation: Separates membrane fractions by buoyant density; suitable for plasma membrane and organelle membranes when starting material is abundant.
- Affinity-based enrichment: Antibody-capture using immobilized antibodies or lectin affinity for subpopulations of surface proteins.
- Surface labelling and capture: Chemical labelling of extracellular domains followed by selective capture enriches cell surface proteins while permitting stringent washing to remove associated cytosolic contaminants.
For extraction and solubilisation:
- Use detergent or organic solvent systems validated to solubilise transmembrane domains while remaining compatible with downstream LC–MS/MS.
- Consider sequential extraction (mild to strong) to fractionate peripheral vs integral membrane proteins.
- When intact-protein analysis is planned, adjust solubilisation to preserve native mass and PTMs.
Protein identification by advanced LC–MS/MS
High-resolution LC–MS/MS is the workhorse for the identification of membrane proteins. Key considerations:
- Peptide-centric (bottom-up) workflows provide broad coverage; hydrophobic peptides often require optimized chromatographic gradients and pre-fractionation.
- Intact-protein (top-down) approaches can resolve proteoforms and post-translational modifications directly, valuable when PTM mapping is a primary objective.
- Instrument platform features such as high resolving power, accurate mass, and sensitive fragmentation methods improve identification confidence for low-abundance membrane proteins.
Quantitative Membrane Proteomics: Strategy Selection
Quantitative analysis is crucial for comparing the abundance of membrane proteins across samples or experimental conditions. Creative Proteomics' Membrane Proteomics services provide flexible quantitative strategies to accommodate various study designs and throughput requirements.
Label-Based Quantification Approaches
Label-based strategies enable accurate relative quantification by incorporating stable isotopes or chemical tags during sample preparation. Use chemical or metabolic labels when you need precise, relative comparisons across a small set of samples (for example, 4–16 conditions). Labels let you run samples together, reducing run-to-run variation and improving fold-change accuracy. Best for discovery experiments where precise control of technical variability is crucial.
Label-Free Quantification Approaches
Label-free strategies rely on signal intensity or targeted monitoring to quantify proteins across samples without chemical labeling. Techniques such as targeted and data-independent acquisition support scalable analysis and are well suited for verification studies or large sample sets. Label-free methods provide flexibility while maintaining quantitative robustness.
- Targeted methods (SRM/MRM, PRM): Allow sensitive, reproducible quantification of preselected peptides; ideal for verification of biomarker candidates.
- Data-independent acquisition (DIA): Provides reproducible, broad coverage across cohorts without labels; suitable for larger-scale profiling where consistent sampling depth is required.
Deliverables and Reporting Standards
- Detailed experimental protocol and QC metrics.
- Raw and processed LC–MS/MS data and identification/quantification tables.
- Annotated lists of membrane proteins with topology and functional annotation.
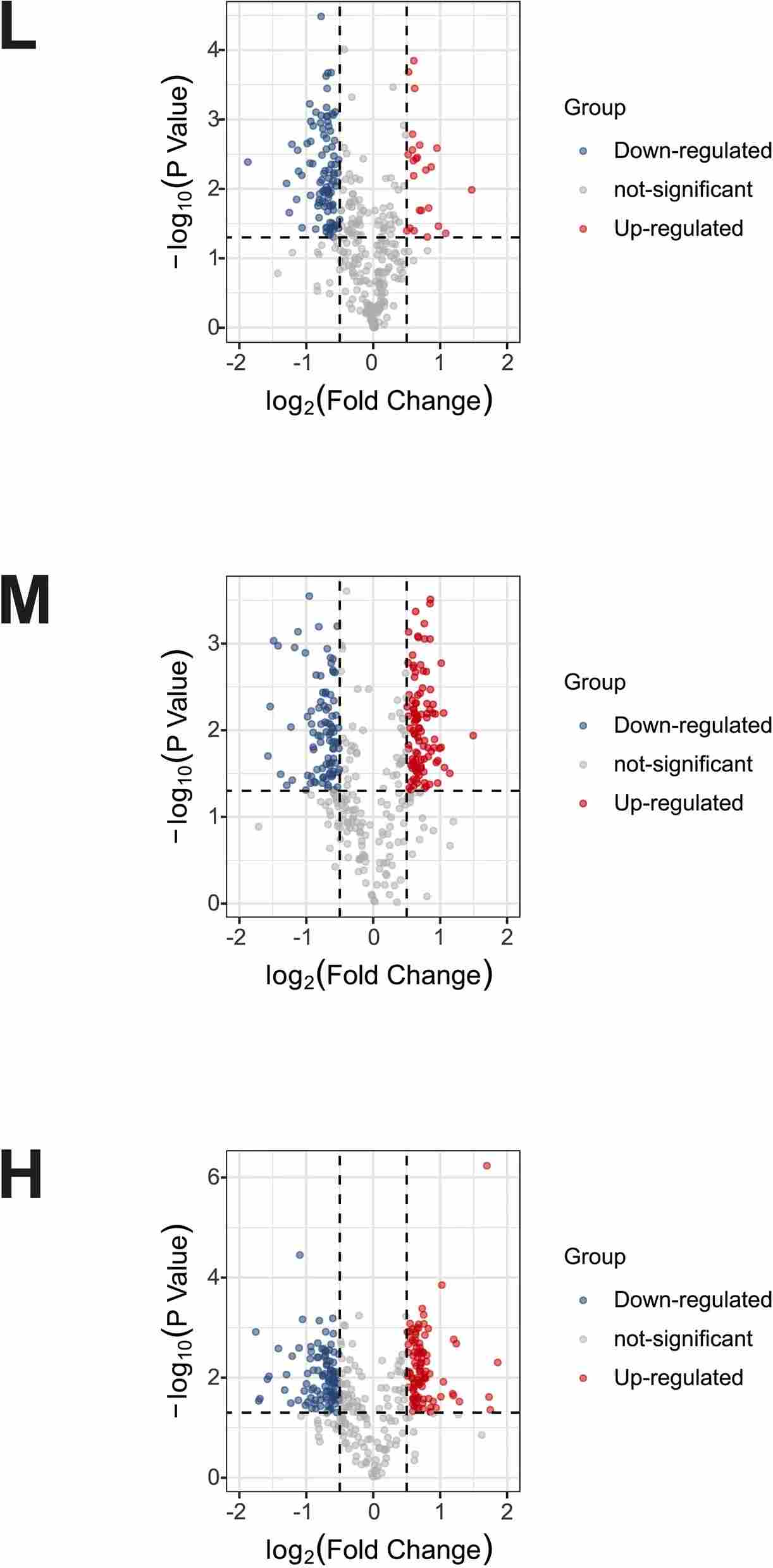
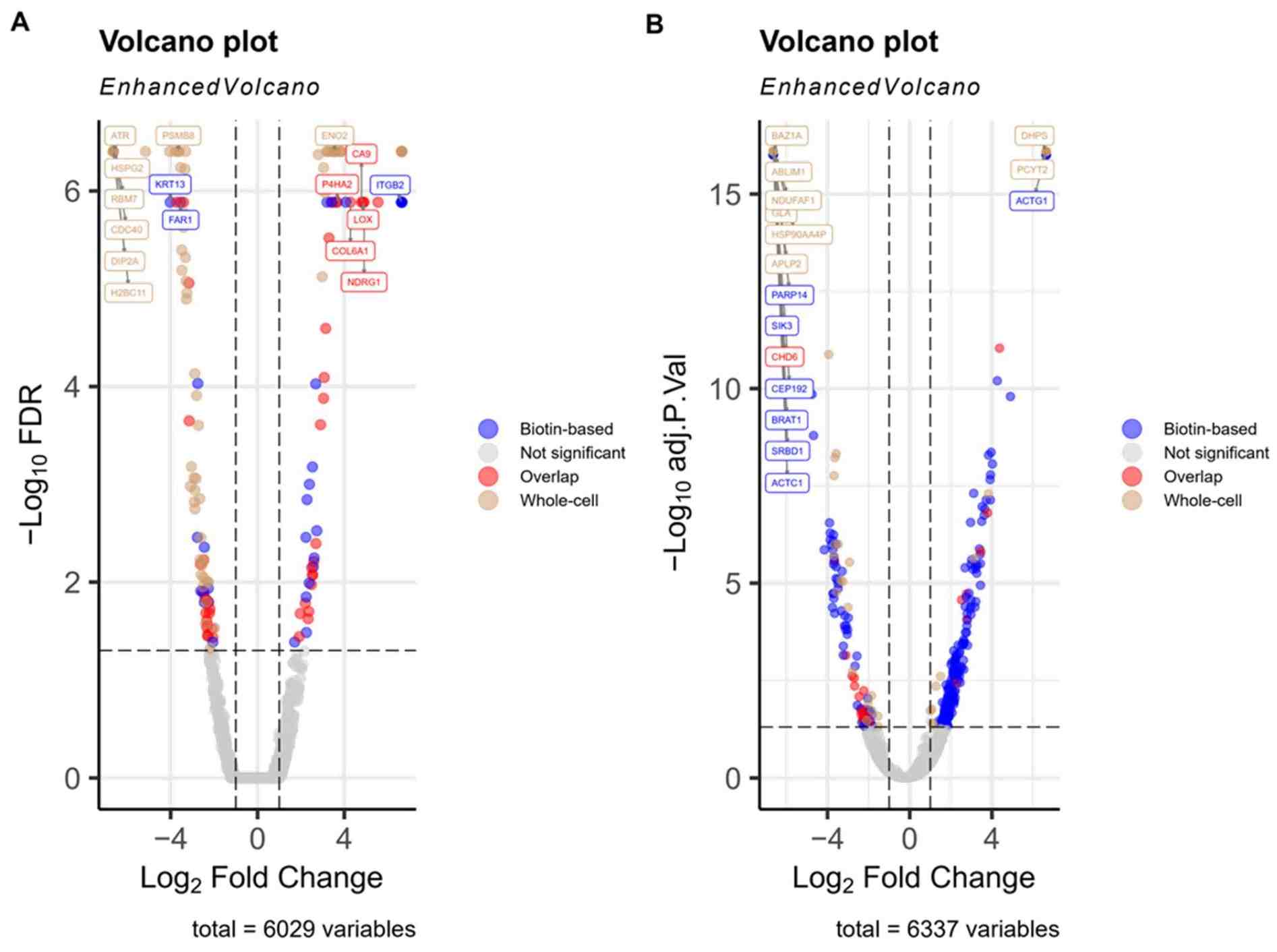
- Publication-ready visualisations (heatmaps, volcano plots, interaction networks) for research dissemination.
Bioinformatics, Topology Prediction, and Functional Interpretation
Bioinformatics turns identifications into biological insight. Essential analysis components include:
- Subcellular localisation and transmembrane topology prediction to confirm membrane assignment and infer orientation.
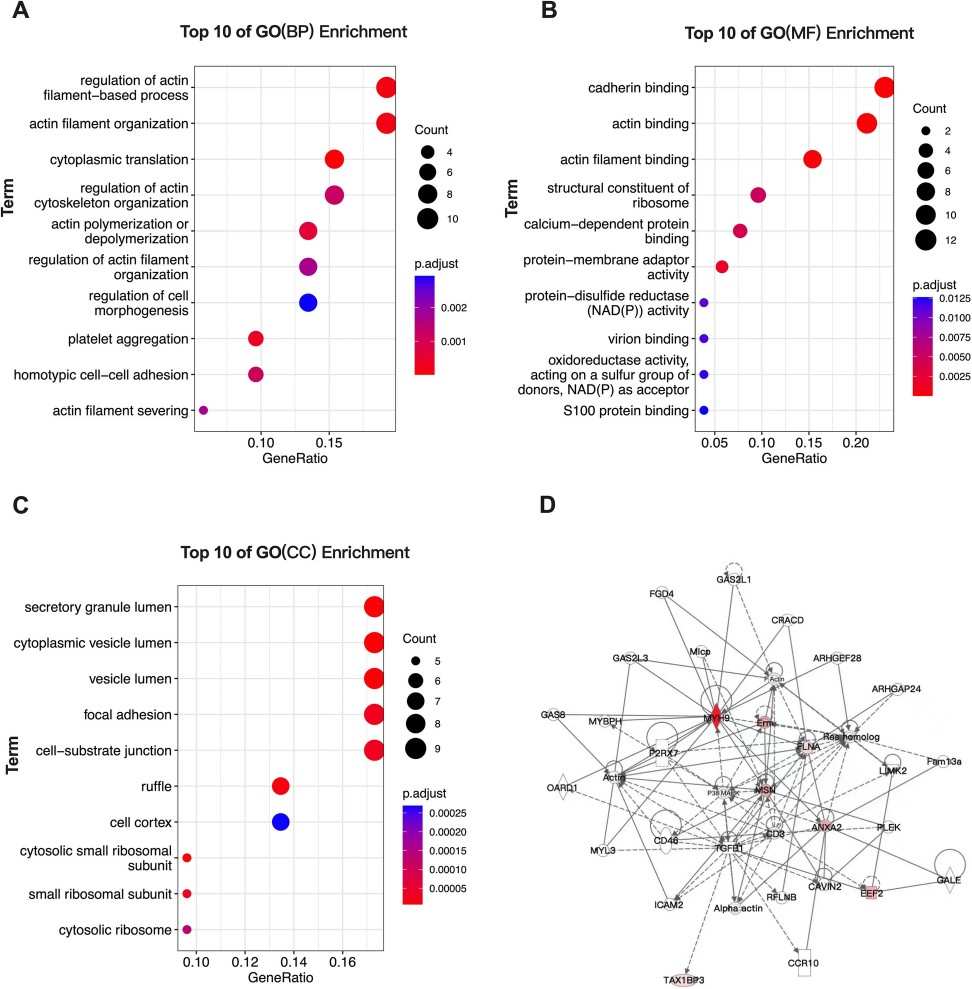
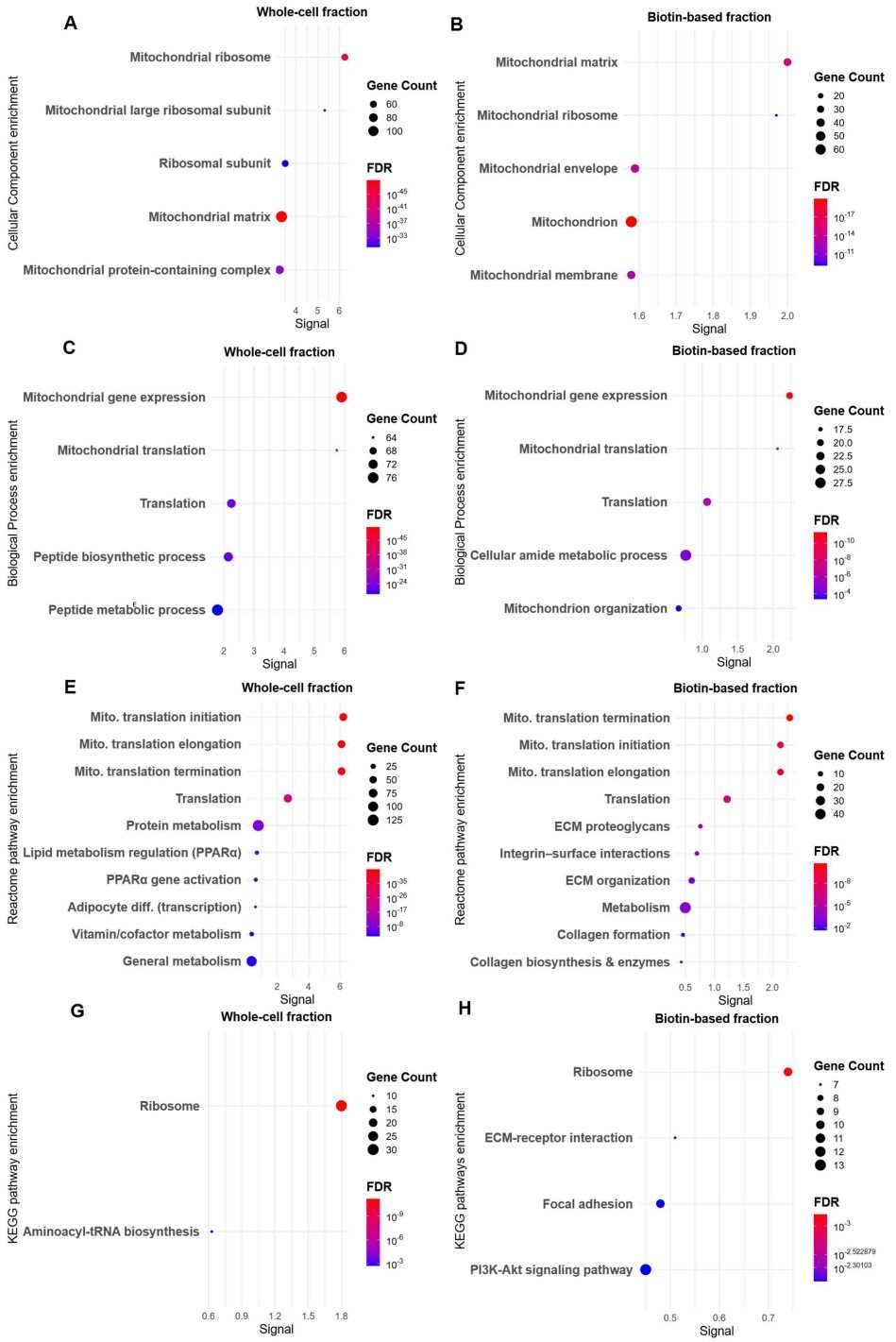
- Gene Ontology (GO) and pathway (e.g., KEGG) enrichment to identify over-represented biological processes.
- PPI networks to prioritise proteins with central roles or potential regulatory connections.
- Proteoform and PTM mapping to detect modification-dependent regulation.
Post-Translational Modification (PTM) Analysis of Membrane Proteins
Membrane proteomics workflows can be adapted to systematically detect and characterize PTMs, including phosphorylation, glycosylation, ubiquitination, and lipidation, which regulate membrane protein localization, stability, and signaling interactions. From a methods perspective, PTM analysis typically begins with optimized membrane enrichment and extraction protocols that preserve labile modifications.
- Phosphorylation: Phosphopeptide enrichment techniques combined with high-resolution LC–MS/MS allow site-specific mapping of low-abundance phosphorylated residues on receptors and transporters.
- Glycosylation: Glycosylation analysis often involves glycopeptide capture, deglycosylation-assisted site mapping, and intact glycopeptide profiling to define site occupancy and glycan heterogeneity.
- Lipid: Lipid modifications, such as acylation or prenylation, are captured using hydrophobic-compatible extraction and MS-compatible enrichment to preserve membrane association.