Introduction to Phosphoproteomics
Phosphoproteomics is a specialized branch of proteomics focused on identifying and quantifying phosphorylated proteins within biological systems. Phosphorylation — the reversible addition of a phosphate to serine, threonine, or tyrosine amino acids — is a key switch that controls protein activity and cellular signaling. Practically, phosphoproteomics uses mass spectrometry together with simple chemical enrichment steps to pull out phosphorylated peptides, so as to detect and quantify phosphorylation across the proteome. This lets researchers map which kinases act on which substrates, track changes in signaling over time or after treatment, and identify location-specific biomarkers or drug targets in diseases such as cancer and neurodegeneration.
What is Spatial Phosphoproteomics?
Protein phosphorylation is one of the most common and essential protein modifications, regulating nearly all cellular signaling processes. Traditional phosphoproteomics measures overall phosphorylation levels but misses where these changes happen within tissues. Spatial phosphoproteomics solves this by combining protein analysis with imaging, revealing how phosphorylation patterns are organized in specific cells and tissue regions. This approach gives researchers a clearer picture of signaling events, disease mechanisms, and potential drug targets in their natural context.
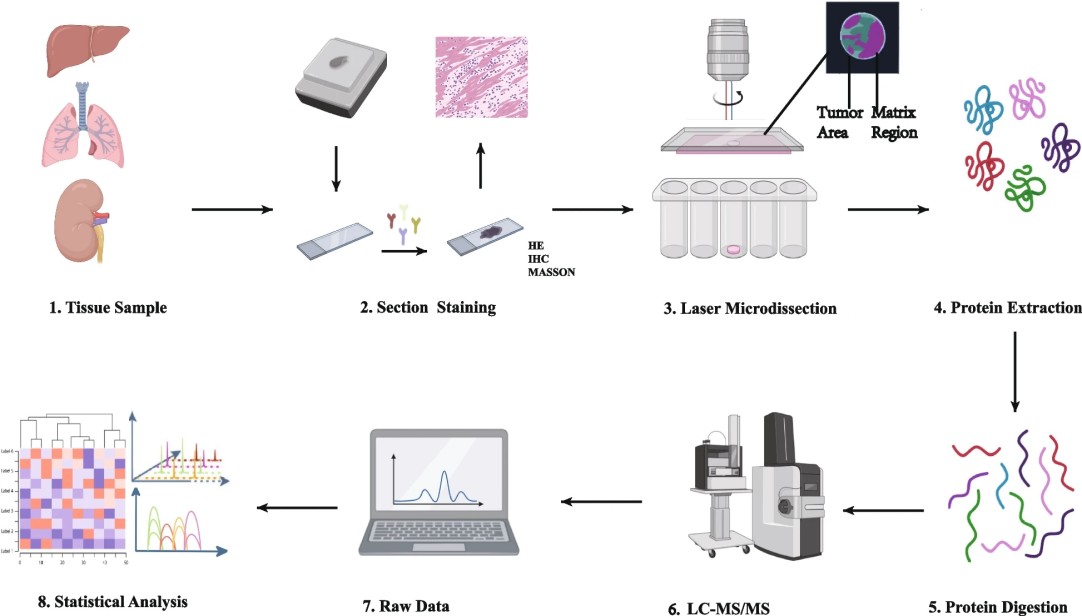
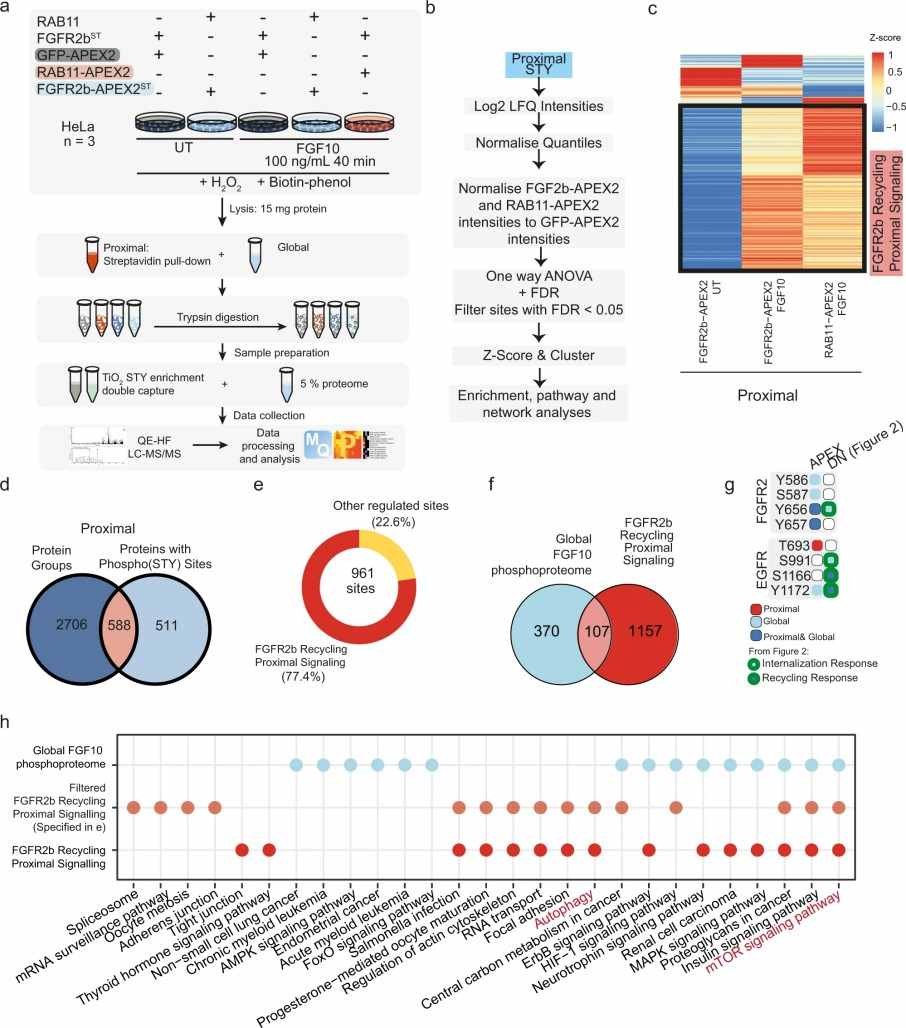
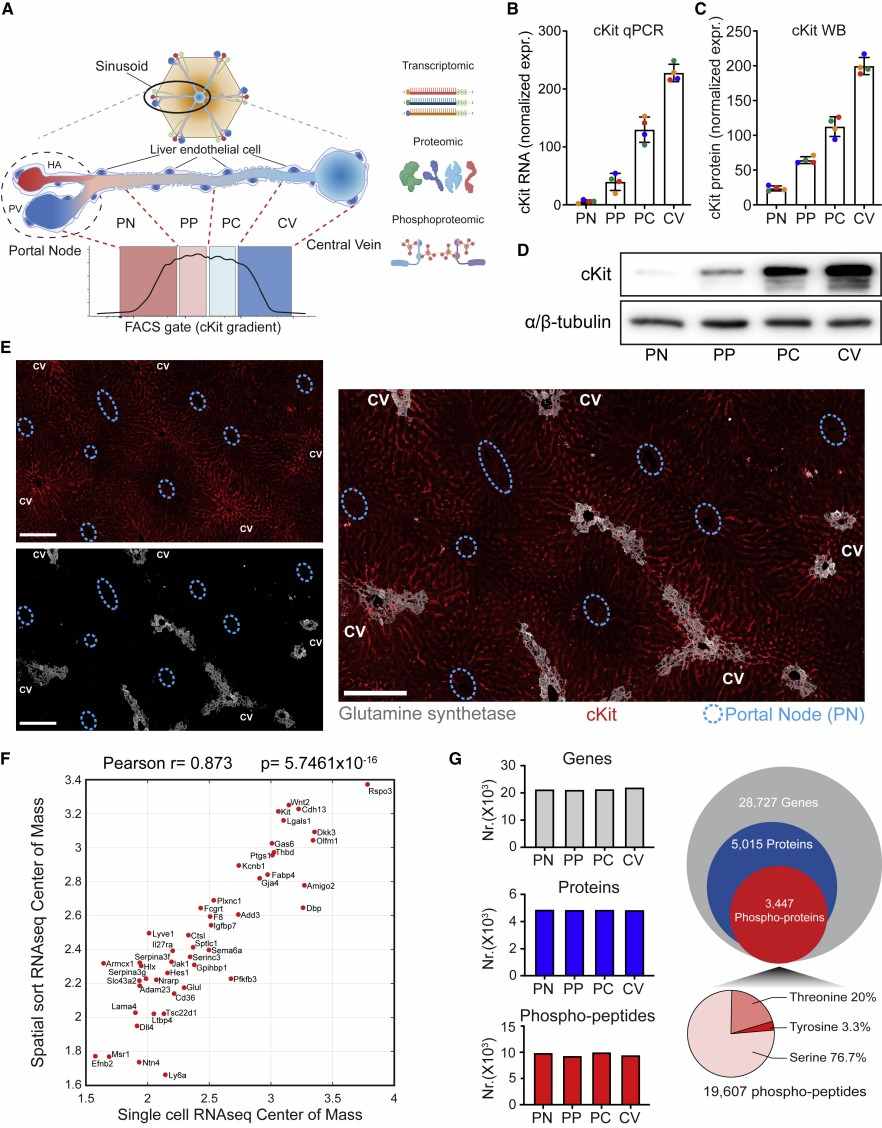
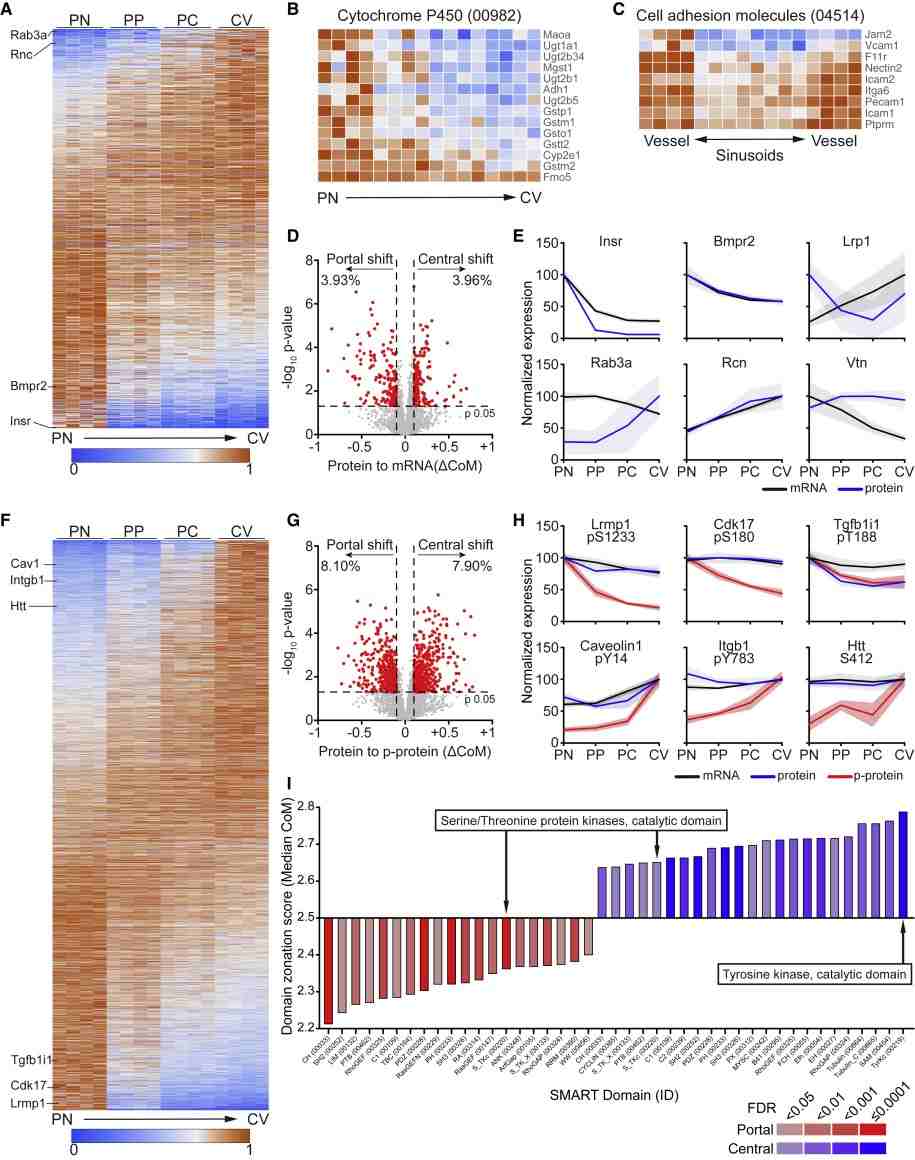
Figure 1. Operational procedure of spatial proteomics mass spectrometry methods (Wu M, et al. 2024).
Why Spatial Context Matters in Phosphoproteomics?
The spatial localization of phosphorylation events is a key determinant of their biological function. Kinase activities, substrate interactions, and signaling cascades occur in highly organized cellular microenvironments. Traditional phosphoproteomics, while comprehensive, averages signals across diverse cell populations, masking critical spatial heterogeneity. Spatial phosphoproteomics overcomes this limitation by analyzing phosphorylated proteins in their native locations within tissues. This allows:
Map signaling gradients within tumors or organs.
Identify phosphorylation hotspots in disease-relevant microenvironments.
Link kinase activity to specific cellular contexts.
Advantages of Creative Proteomics' Spatial Phosphoproteomics Service
- Multi-Sample Compatibility and High Flexibility: Our spatial workflow supports multiple sample formats (FFPE, OCT, frozen) and staining methods (HE, IHC, IF). While spatial phosphoproteomics currently focuses on frozen sections, we ensure compatibility across different experimental designs.
- Ultra-Sensitive and Deep Detection: With advanced mass spectrometry systems and optimized sample preparation, we achieve high detection depth and precision even from nanogram-level protein inputs. Our workflow delivers consistent phosphoproteomic coverage exceeding 5000+ phosphosites per analysis.
- Exceptional Reproducibility and Reliability: Technical replicates exhibit Pearson correlation coefficients above 0.95, ensuring robustness of results across biological and technical repeats.
- Integrated Infrastructure and Expertise: Our facility houses state-of-the-art instruments—including cryostats, LCM, and digital slide scanners—enabling a seamless, one-stop spatial proteomics solution from imaging to MS detection.