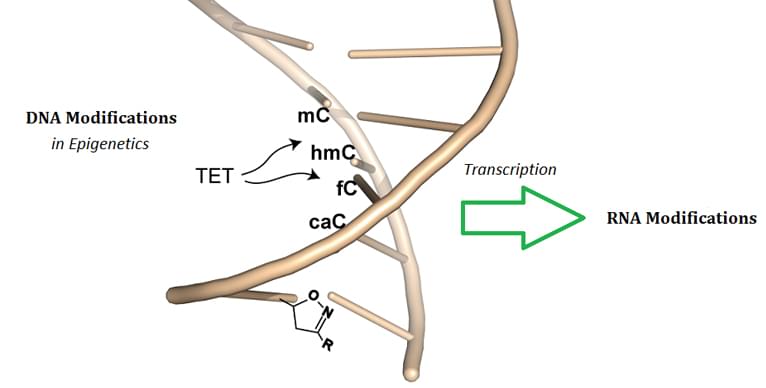
DNA methylation is a process by which the methyl groups are covalently transferred to cytosine & adenine of DNA nucleotides (adenine methylation is restricted to prokaryotes), by the catalysis of DNA methyltransferases. Typically acting to suppress gene transcription, DNA methylation is essential for normal development and associated with a series of key processes and biological activities, including genomic imprinting, X-chromosome inactivation, suppression of repetitive elements, and carcinogenesis. In mammals, insects, bacteria, fungi and plants, DNA methylation is widely detected in the species above; DNA methylation typically occurs at the 5 position of cytosine in a CpG dinucleotide contect, which can selectively reduce specific gene expression; non-CpG methylation is prevalent in embryonic stem cells, and has also been indicated in neural development.
As one of the major epigenetic processes, it is widely accepted that there is an underlying relationship between DNA methylation, chromatin structure & human malignancies. Because of the significance of DNA methylation as biomarkers in cancer molecular diagnostics, aging, atherosclerosis and other diseases, currently several analytic have been developed likely chemical assay or biochips. Now with the development of mass spectrometry, LC-MS/MS can work as a sensitive and reliable analytic method to globally profile methylation with high throughput, ultra-sensitivity and high confidence. Oligonucleotides containing unusual modifications can therefore be verified by mass spectrometery rapidly. And the typical work-flow for LC-MS based analysis would be as follow:
- HPLC can be used to separate the different deoxyribonucleotide fragments in a base-specific digested sample of DNA, yielding different retention times on the chromatographic column;
- In tandem mass spectrometry, the protonated deoxyribonucleotides are sequentially selected by the first mass analyzer, and fragmented in the collision cell to generate product ions;
- The patterns of fragment ions from the selected precursor would be scanned by the second mass analyzer. The bioinformatic tools would be applied for identifications of DNA sequence fragments and DNA methylation.
Creative Proteomics have developed valid SOPs of DNA isolation and digestion, HPLC separation, and analysis with tandem mass spectrometry, based on the reliable protocols from reputable journals. Operating the advanced and integrated facilities, our bioanalytical staff can generate robust analytic results for your project.





